True or False: Genetics only takes place at the DNA level.
False
- Genetics takes place on several levels, including organismal, cellular, chromosomal, and molecular
Organismal Genetics
Observes heredity of the whole organism
Cellular Genetics
Looks at single cells
Chromosomal Genetics
Examines the characteristics and actions of chromosomes
Molecular Genetics
Deals with biochemistry of genes
_______ is the sum total of genetic material of an organism.
Genome is the sum total of genetic material of an organism.
Most of the genome exists in the form of ________ while some may appear in ______________ forms (plasmids, organelles)
Most of the genome exists in the form of chromosomes while some may appear in nonchromosomal forms (plasmids, organelles)
______ tiny extra pieces of DNA
Plasmids
What two organelles have their own DNA?
- Mitochondria
- Chloroplast
Genomes of cells are made of ____, but viruses contain ______ or _____.
Genomes of cells are made of DNA, but viruses contain either DNA or RNA.
The study of an organism's entire genome is known as...
Genomics
_______ is a distinct cellular structure composed of a neatly packaged DNA molecule.
Chromosome
How do eukaryotic and bacterial chromosome differ?
- The structure of eukaryotic chromosomes consists of DNA tightly wound around histone proteins, whereas a bacterial chromosome is condensed and jammed into a packet form by means of histonelike proteins.
- Eukaryotic chromosomes are located in the nucleus; they vary in number from a few to hundreds; they can occur in pairs (diploid) or singles (haploid); and they have a linear appearance.
- Most bacteria have a single, circular (double-stranded) chromosome, although many bacteria have multiple, circular chromosomes and some have linear chromosomes.
____ site on the chromosome that provides basic information for a certain cell function.
Gene
Genes fall into three categories:
- Structural genes that code for protein
- Genes that code for the RNA machinery used in protein production
- Regulatory genes that control gene expression
_________ is the sum of all types of genes constituting an organism's distinctive genetic makeup.
Genotype is the sum of all types of genes constituting an organism's distinctive genetic makeup.
_______ is the expression of the genotype that creates certain structures or functions (traits).
Phenotype is the expression of the genotype that creates certain structures or functions (traits).
True or False:
All organisms contain more genes in their phenotype than are manifested as a genotype at any given time.
False
All organisms contain more genes in their genotypes than are manifested as a phenotype at any given time
What does this statement mean?
- All organisms contain more genes in their genotypes than are manifested as a phenotype at any given time
The phenotype can change depending on which genes are "turned on" (expressed)
The basic unit of a DNA is....
Nucleotide
A nucleotide consists of:
- Phosphate
- Deoxyribose sugar
- Nitrogenous base
What are the two types of nitrogenous bases?
- Purines (A, G)
- Pyrimidines (C, T)
- Adenine (A) always pairs with Thymine (T)
- Guanine (G) always pairs with Cytosine (C)
What is an important characteristic of DNA?
DNA is arranged antiparallel
- One side of the helix runs in the opposite direction of the other: 5′ to 3′ in one direction and 3′ to 5′ in the other direction
- Significant factor in DNA synthesis and protein production
Two significances of the antiparallel DNA structure
- Maintaining the code during reproduction - Each strand provides a template for the replication of a new molecule guaranteeing accurate duplication
- Providing variety - The precise sequence of bases constitutes a genetic program responsible for the unique qualities of each organism
Most DNA is replicated in a _____-_________ manner, meaning that one strand will always be the original strand and the other strand will be the newly synthesized strand.
Semi-Conservative
Steps of the replication (simplified)
- Circular DNA has a special origin site where replication begins. When strands are separated, two replication forks form, and a DNA polymerase III complex enters at each fork. The DNA polymerases proceed in both directions along the DNA molecule, attaching the correct nucleotides according to the pattern of the template.
Helicase
Unzipping the DNA helix
Primase
Synthesizing RNA primer
DNA Pol III
Adding bases to the new DNA chain and proofreading the chain for mistakes
DNA Pol I
Removing primer, closing gaps, and repairing mismatches
Ligase
Final binding of nicks in DNA during synthesis and repair
Topoisomerase I
Making single-stranded DNA breaks to relieve supercoiling at origin
Topoisomerase II (DNA gyrase) and 4
Making double-stranded DNA breaks to remove supercoiling ahead of origin and separate replicated daughter DNA molecules
In the semi-conservative model...
- Each daughter molecule is identical to the parent in composition
- Neither is completely new
- The template strand is an original parental DNA strand
True or False: DNA pol III can only add nucleotides in 3' to 5' direction.
False
- DNA pol III can only add nucleotides in 5' to 3' direction
Functions of DNA pol lll:
- Functions once the DNA helix strands are unwound and separated
- Synthesizes a new daughter strand of DNA using the parental strand as a template
- Can only add nucleotides to an existing chain—cannot begin synthesizing a chain of nucleotides
- Can only add nucleotides in the 5′ to 3′ direction
Replication Fork
- The place in the helix where the strands are unwound and replication is taking place
- Each circular DNA molecule will have two replication forks
Primer
- A length of RNA that is inserted initially during replication before being replaced by DNA
- This RNA is eventually replaced by DNA after replication.
True or False: Both eukaryotes and prokaryotes have replication forks.
True
- Eukaryotes have multiple sites where replication forks are
- Prokaryotes only have one site
The strand of new DNA that is synthesized continuously in a 5' to 3' direction
Leading Strand
The strand of new DNA that must be synthesized in short segments (in a 5' to 3' direction)
Lagging Strand
- This strand is later sealed together to form a strand in the 3' to 5' direction
What are Okazaki fragments?
Short segments of DNA synthesized in a 5' to 3' direction which are then sealing together to form the 3' to 5' strand
DNA Replication Steps
1. During replication topoisomerases unwind the DNA helix, giving access to helicase (unzipping enzymes) to bind to the dsDNA at the origin. Helicases break the hydrogen bonds holding the two strands together, resulting in two separate strand.
2. Single-stranded binding proteins keep the strand apart.
3. DNA pol III adds nucleotides in accordance with teh template pattern. Note that RNA primase will have already added a short length of RNA. Because DNA polymerase is correctly oriented for synthesis only in the 5' to 3' direction of the new molecule strand, only one strand, called the leading strand, can be synthesized as a continuous, complete strand. The strand with the opposite orientation (3' to 5') is the lagging strand.
- On the lagging strand, polymerase adds nucleotides a few at a time in the direction away from the fork (5' to 3'). As the fork opens up a bit, the next segment is synthesized backward to the point of the previous segment, a process repeated until synthesis is complete.
- In this way, DNA polymerase is able to synthesize the two new strands simultaneously.
____ begin linking fragments, and ___________ causes a double-stranded DNA break separating the two fully replicated daughter molecules.
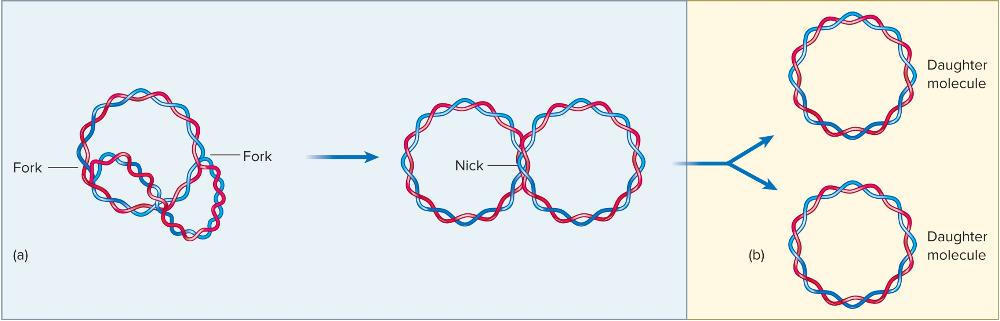
Ligases begin linking fragments, and Topoisomerase IV causes a double-stranded DNA break separating the two fully replicated daughter molecules.
How is replication of eukaryotic DNA similar to replication of bacterial and archaeal DNA?
- Uses a variety of DNA polymerases
- Replication proceeds in both directions but from multiple origins
- Topoisomerases are used to relieve tension in the strand and to recompact the molecule when it is fully replicate
Due to the unidirectional action of DNA polymerase, the 3' end of linear DNA molecules, called ______, cannot be copied entirely
Due to the unidirectional action of DNA polymerase, the 3' end of linear DNA molecules, called telomeres, cannot be copied entirely
_____________: DNA is used to synthesize RNA
_____________: RNA used to produce proteins
Transcription: DNA is used to synthesize RNA
Translation: RNA used to produce proteins
Why is the central dogma incomplete?
- A wide variety of RNAs are used to regulate gene function
- Many genetic malfunctions that cause human disease are found in regulatory RNA, not in genes for proteins
- The DNA that codes for these crucial RNA molecules was once called “junk” DNA
What is the central dogma?
DNA --> RNA --> Protein
Describe the flow of genetic information in microbes.
- DNA is the ultimate storehouse and distributor of genetic information
- DNA must be deciphered into usable language. It does this by transcribing its code into RNA helper molecules that translate that code into protein
- Other sections of the DNA produce very important RNA molecules that regulate genes and their products
What is the connection between gene and protein? In other words, what is the connection between DNA and an organism's trait?
- A protein's primary structure determine its characteristic shape and function
- Proteins ultimately determine phenotype (Proteomics is the study of an organism's complete set of expressed proteins)
- DNA is mainly a blueprint that tells the cell which kinds of proteins to make and how to make them
The DNA molecule is a continuous chain of base pairs, but the sequence must be interpreted in groups of ____ base pairs (a ____). Each _____ as copied into ____ codons will translate into one amino acid.
The DNA molecule is a continuous chain of base pairs, but the sequence must be interpreted in groups of three base pairs (a triplet). Each triplet as copied into mRNA codons will translate into one amino acid.
What are the major participants in transcription and translation?
- mRNA (messenger RNA) - Creator of proteins
- tRNA (transfer RNA) - Adds the amino acids that build your proteins. Coded from mRNA codons
- rRNA (regulatory RNA) - Ribosomal RNA that translates the mRNA using tRNAs
- Ribosomes
- Several types of enzymes
- Many raw material
What is the only type of RNA that is translated?
mRNA (Messenger RNA)
Description and Function of mRNA
- Description - Sequence of amino acids in protein
- Function - Transports DNA master code to the ribosome
Description and Function of tRNA (Transfer RNA)
- Description - A cloverleaf tRNA to carry amino acids
- Function - Brings amino acids to ribosome during translation
Description and Function of rRNA (Ribosomal RNA)
- Description - Several large structural rRNA molecules
- Function - Forms the major part of a ribosome and participates in protein synthesis
Description and Function of Micro (miRNA), antisense, riboswitch, and small interfering (siRNA)
- Description - Regulatory RNA
- Function - Regulation of gene expression and coiling of chromatin
Description and Function of Primer
- Description - An RNA that can begin DNA replication
- Function - Primes DNA
Description and Function of Ribozymes and Spliceosomes (snRNA)
- Description - RNA enzymes, parts of splicer enzymes
- Function - Remove introns form other RNAs in eukaryotes
How are RNAs different from DNA?
- Single stranded molecules that exists in helical form (can assume secondary and tertiary levels of complexity)
- Contains uracil (U) instead of thymine as the complementary base pair to adenine
- Sugar is ribose rather than deoxyribose
Messenger RNA (mRNA)
- A transcript of a structural gene or genes in the DNA
- Synthesized in a process similar to synthesis of the leading strand during DNA replication
_____ are a series of triplet bases that hold the message of the transcribed mRNA that will be translated into an amino acid.
Codons are a series of triplet bases that hold the message of the transcribed mRNA that will be translated into an amino acid.
tRNA are known as the _______ molecule.
Adapter
Which type of RNA contains sequences of bases that form hydrogen bonds with complementary sections of the same tRNA strand?
Transfer RNA (tRNA)
Where is the anticodon found?
- Found at the bottom loop of the cloverleaf
- Designates the specificity of the tRNA and complements the mRNA codon
What is the function of rRNA?
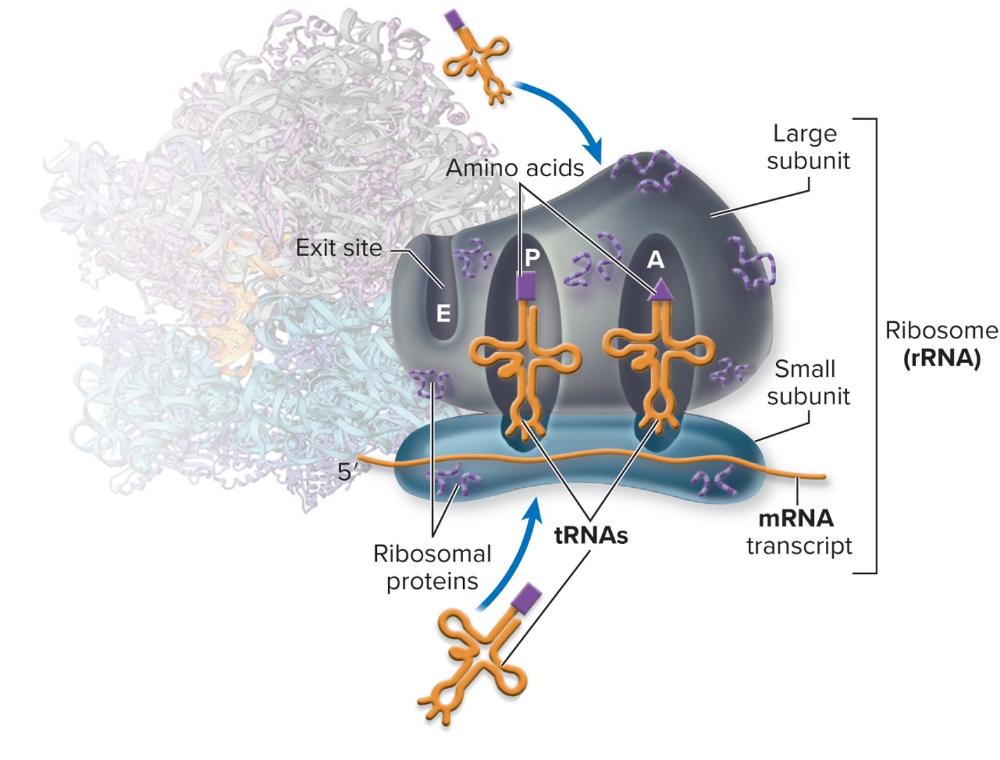
- A long RNA molecule and proteins form complex three-dimensional shapes to create the two subunits of the ribosome
- A metabolically active bacterial cell can contain up to 20,000 70S ribosomal units
- The two subunits engage in the final translation of the genetic code
- Basically the large subunit and the small subunit come together to for the rRNA
Three steps of transcription are:
- Initiation
- Elongation
- Termination
When is transcription initiated?
- Transcription is initiated when RNA polymerase recognizes a segment of the DNA called the promoter region.
- This region consists of two sequences of DNA just prior to the beginning of the gene to be transcribed. These promoter sequences provide the signal for RNA polymerase to bind to the DNA. As the DNA helix unwinds, the polymerase first pulls the early parts of the DNA into itself, a process called "DNA scrunching," and then, having acquired energy from the scrunching process, begins to advance down the DNA strand to continue synthesizing an RNA molecule complementary to the template strand DNA. The nucleotide sequence of promoters differs only slightly from gene to gene, with all promoters being rich in adenine and thymine.
- Only one strand of DNA, called the template strand, is copied by RNA polymerase.
What happens during the elongation step of transcription?
- During elongation, which proceeds in the 5' to 3' direction (with regard to the growing RNA molecule), the mRNA is assembled by the addition of nucleotides that are complementary to the DNA template.
- Remember that uracil is placed as adenine's complement.
- As elongation continues, the part of DNA already transcribed is rewound into its original helical form
What happens during the termination step of transcription?
- RNA polymerase continues to make its way down the template strand until it reaches a specific sequence that signals the end of the transcript.
- There is more than one mechanism for ending the process, but it is always signaled by the DNA sequence in the template strand.
After the mRNA is made from _______, the mRNA is now ready to be _________.
After the mRNA is made from transcription, the mRNA is now ready to be translated.
Central principle of translation:
- mRNA nucleotides are read in codons or groups of three
- The codon dictates which amino acids are added to the growing chain
- Except for a very few cases, this code is universal for bacteria, archaea, eukaryotes, or viruses
In prokaryotes, the start codon is...
AUG which is f-methionine
What are the three stop codons?
UGA, UAG, UAA
The master genetic code is:
- Redundancy
- Wobble
What is meant by the redundancy concept of codons?
- Certain amino acids are represented by multiple codons
- Allows for the insertion of correct amino acids even when mistakes occur in the DNA sequence
What is meant by the wobble hypothesis?
- Only the first two nucleotides are required to encode the correct amino acid
- The third nucleotide does not change its sense
- Permits some variation or mutation without altering the message
What are the three stages of translation?
- Initiation
- Elongation
- Termination
What are all the elements needed to synthesize a protein?
- mRNA
- tRNA
- Amino Acids
- Ribosomes
Start Codon
- The first three RNA nucleotides that signal the beginning of the message
- Always A U G
Stop Codon
- Nonsense codons: one of three codons that has no corresponding tRNA and causes translation to be terminated
- U A A, U A G, and U G A
Translocation
- The process of shifting the ribosome down the mRNA strand to read new codons
Can tRNA bind without the presence of ribosome?
NO
Brief Steps of Translation
1. Entrance of tRNAs 1 and 2
2. Formation of peptide bond
3. Discharge of tRNA 1 at E site
4. First translocation
5. Formation of peptide bond
6. Discharge of tRNA 2; second translocation; enter tRNA 4
7. Formation of peptide bond
A little more in depth of translation:
- Initiation – The ribosome assembles and binds the 5’ end of the mRNA. Once it hits the start codon, it can now begin to recruit tRNAs associated with the codons marking initiation. There are three sites: EPA, for a tRNA entrance, peptide bonding, and exit. Note that bacteria have the F-methionine, which is usually cleaved afterwards.
- After initiation is elongation. This begins with the second tRNA entering the A site. The amino acid attached to both tRNAs form a peptide bond and the F-methaionine, or basically that tRNA in that P site, is transferred to the A site and the corresponding tRNA in that A site. Then, the translocation of the ribosome complex moves the empty tRNA from the P to the E site. Now the tRNA with two amino acid moves to P site. Now, A site to open and another amino acid can enter.
- At the end, a special enzyme, not a tRNA, comes in and cleaves the peptide chain from the tRNA in the P site.
Explain the entirety of translation
1. The mRNA molecule leaves the DNA transcription site and is transported to ribosomes in the cytoplasm. Ribosomal subunits come together and form sites to hold the mRNA and tRNAs. The ribosome begins to scan the mRNA by moving in the 5' to 3' direction along the mRNA. The first codon it encounters called the start codon. With the mRNA message in place on the assembled ribosome, the next step in translation involves entrance of tRNAs with their amino acids. The pool of cytoplasm contains a complete array of tRNAs, previously charged by having the correct amino acid attached. The step in which the complementary tRNA meets the mRNA code is guided by the two sites on the large subunit of the ribosome called the P site and the A site. The ribosome also has an exit or E site where used tRNAs are released.
2. Rules of pairing dictate that the anticodon of tRNA must be complementary to the mRNA codon. The ribosome shifts its "reading frame" to the right along the mRNA from one codon to the next. This brings the next codon into place on the ribosome and makes a place for the next tRNA to enter the A position. A peptide bond is formed between the amino acids on the adjacent tRNAs, and the polypeptide grows.
Elongation beings with the filling of the A site by a second tRNA. The identify of this tRNA and its amino acid is dictated by the second mRNA codon.
3. The entry of tRNA into the A site brings two adjacent tRNAs in favorable proximity for a peptide bond to form between the amino acids they carry.
For the next step to proceed, some room must be made on the ribosome, and the next codon in sequence must be brought into position for reading. This process is accomplished by translocation, the enzyme-directed shifting of the ribosome to the right along the mRNA strand, which causes the blank tRNA 1 to be discharged from the ribosome at the E site.
4. Dipeptide moves to P site. Site A is empty. tRNA that has been released is now free to drift off into the cytoplasm and become recharged with an amino acid for later additions to this or another protein.
The stage is now set for the insertion of tRNA 3 at site A as directed by the third mRNA codon. This insertion is followed once again by peptide bond formation between the dipeptide and amino acid 3 (making a tripeptide), splitting of the peptide from tRNA 2, and translocation.
5. Releases tRNA 2, shifts mRNA to next position, moves tRNA 3 to position P, and opens position A for next tRNA.
6. From this point on, peptide elongation proceeds. 7. The termination is brought by presence of at least one special codon occurring just after the codon for the last amino acid. When this codon is reached, a special enzyme breaks the bond between the final tRNA and the finished polypeptide chain, releasing it from the ribosome.
What happens after protein synthesis?
Posttransitional modifications
- Proteins begin to fold upon themselves to achieve their tertiary conformation even before the peptide chain is released
- Formyl methionine may be clipped off
- Cofactors may be added
- Some join with other proteins to form a quaternary structure, meaning that there are multiple proteins making up one functional enzyme
Why is protein assembly line in bacteria faster?
- There are multiple ribosomes attached to one mRNA which speeds up the process of protein production, making it a much more efficient process.
- The mRNA transcript encounters ribosomal parts immediately as it leaves the DNA.
- The ribosomal factories assemble along the mRNA in a chain, each ribosome reading the message and translating it into protein. Many products will thus be well along the synthetic pathway before transcription has even terminated.
COVID-19 viruses have...
- Contains a viral mRNA that codes for the surface protein of the virus
How does the COVID-19 mRNA vaccine work?
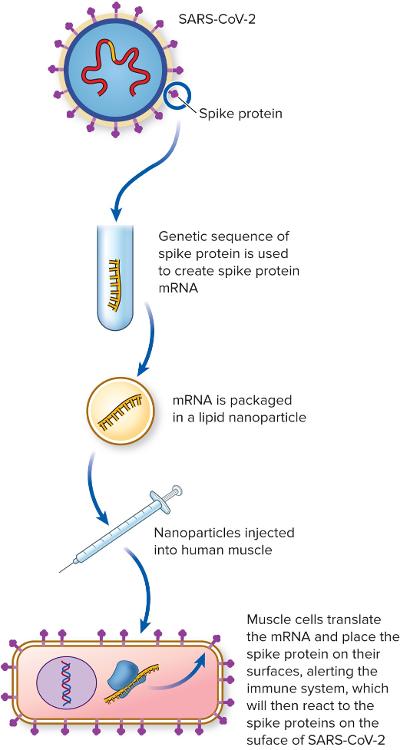
- Once injected into human arms, muscle cells use their translation machinery to create and place on their cell surface the viral protein
- Normal immune system screening processes detect it and build specific immune defenses against that protein
What are differences between prokaryotic and eukaryotic transcription and translation?
- Co-transcriptional translation only occurs in bacteria and archeae
- A U G codes for a different form of methionine in eukaryotes
- Eukaryotic mRNA’s only code for one protein; bacterial mRNA’s often contain information from several genes in a series
____ are the coding regions whereas the _____ are intervening sequences of bases that do not code for protein.
Exons are the coding regions whereas the introns are intervening sequences of bases that do not code for protein.
What recognizes the exon-intron junctions and enzymatically cuts them?
Spliceosome
- Loops introns into lariat-shaped pieces, excises them, and joins the exons end to end
- Completed mRNA strand can then proceed to the cytoplasm for translation
Operons
- A coordinated set of genes regulated as a single unit
- Found only in bacteria and archaea
- Can be inducible or repressible, determined by how transcription is affected by the environment surrounding the cell
Inducible Operons
- Contain genes that make proteins that help metabolize nutrients
- Operon is turned on (induced) by the nutrient for which the structural genes encode
- Enzymes needed to metabolize a nutrient are only produced when that nutrient is present in the environment
- Naturally turned off and only turned on when those nutrients are available in the environment
Repressible Operons
- Contain genes coding for enzymes that synthesize amino acids
- Their normal state is “on”
- When there is enough product synthesized by the enzyme the operon is turned off (repressed)
What type of operon contains genes that make proteins that help metabolize nutrients?
Inducible Operons
What type of operon contain genes that synthesize amino acids
Repressible
Three features of the lac operon:
- Regulator
- Control Locus
- Structural Locus
Regulator
Composed of the gene that codes for the repressor, a protein capable of repressing the operon
Control Locus
- Promoter: recognized by RNA polymerase
- Operator: acts as an on/off switch for transcription
Structural Locus
Made up of three genes, each coding for a different enzyme needed to catabolize lactose
What is the main purpose of the lactose operon?
- The main purpose of this system is to switch from glucose metabolism to lactose once glucose is not available.
- This operon has the regulator, which makes the repressor protein, and that repressor protein binds to the operator, and has the control locus, which contains the promotor lac-P and the operator lac-O portions of the operon.
- Three genes that are needed to make sure the bacteria can use lactose are called collectively the structural genes or the structural locus.
- Normally, it will use glucose when glucose is present, however, if there is no glucose and high levels of lactose, this system will be turned on.
The repressor protein for the lactose operon is...
Allosteric
What does allosteric protein do in a lactose operon?
It is the repressor protein:
- Two binding sites: one for the operator sequence on the DNA and another for lactose
- In the absence of lactose, the repressor binds to the operator, blocking transcription of structural genes
- The regulator gene lies upstream of the operator and is transcribed constitutively
- If lactose is bound and available, then the lactose portion is bound and the operator sequence is free.
- Repressor protein is usually always active and blocks RNA polymerase from being able to transcribe that operon.
Describe the lac operon in bacteria
1. This operon is normally in an "off" mode and does not initiate transcription when the appropriate substrate is absent.
2. If lactose is added to the cell's environment, it triggers events that turn the operon on.
3. The structural genes are transcribed in a single unbroken transcript coding for all three enzymes. (During translation, however, each protein is synthesized separately).
The lac operon is usually... inducible.
- So, the lactose or the lac operon in bacteria, you have the regulator that produces the repressor protein (normally, this repressor protein is always bound to a portion of operator, blocking RNA polymerase from being able to transcribe and translate the structural genes). However, in the absence of glucose and presence of lactose, lactose induces and binds to the repressor protein, causing a conformational change, where now the repressor protein no longer binds to the operator, leaving it free and open for RNA polymerase to bind and become active (and being able to transcribe and translate the structural proteins).
- If that lactose is then depleted and there’s no more binding, then the system returns back to normal where it is now being repressed
Repressible operon
- Repressible operons are usually in the on mode
- Will only be turned off only when the nutrient is no longer required
- Excess nutrient serves as a corepressor to block the action of the operon
Repressible Operon: Control of a Gene Through Excess Nutrients
- Operon on. A repressible operon remains on when its nutrients products are in great demand by the cell because the repressor is unable to bind to the operator at low nutrient levels
- Operon off. The operon is repressed when nutrient builds up, serving as a corepressor, activates the repressor. The repressor complex affixes to the operator and blocks RNA polymerase and further transcription of genes for nutrient synthesis.
What two ways do eukaryote cells regulate genes?
- Transcription Factors
- DNA "knot"
Transcription Factors
- Insert into the grooves of the DNA molecule and enhance the transcription of specific genes
- Regulate gene expression in response to environmental stimuli
DNA "knot"
- Cytosines bind to other cytosines forming a “knot” in the helix of DNA
- Blocks the promoter region of genes in order to stop transcription
Phase Variation
The result of bacteria turning on or off a complement of genes that leads to phenotypic changes
- Heritable—passed down to subsequent generations
- Involves turning on genes mediated by regulatory proteins, as described with operons
What are the drugs the interfere with ribosome?
- Erythromycin
- Spectinomycin
What are the drugs that inhibit protein synthesis?
- Rifamycin
- Actinomycin D
An event in which one bacterium donates DNA to another bacterium.
Recombination
What is the result of recombination?
The end result is a strain different from both the donor and the recipient strain
What is the extrachromosomal DNA adept at moving between cells called?
Plasmids
Any organism that contains and expresses genes that originated in another organism is known as...
Recombinant
What is horizontal gene transfer?
Any transfer of DNA that results in organisms acquiring new genes that did not come directly from parent organisms
Horizontal gene transfer does not include chromosomes. It is about ______.
Plasmids
Can eukaryotic organisms also engage in horizontal gene transfer?
Eukaryotic organisms also engage in horizontal gene transfer, often aided by microbes such as viruses
What are the three types of horizontal gene transfer?
- Conjugation
- Transformation
- Transduction
Small, circular pieces of DNA that can replication independently of the bacterial chromosome are...
Plasmids
- Allow transfer of DNA between cells
- Found in most bacteria and in some fungi
- Contain at most a few dozen genes
- Not necessary for survival, but often carry useful traits
Conjugation
- Bacterial "sex"
- Involves a pilli
- Must have a + and - bacteria
Transformation
- "scavenger"
- Free DNA is taken up
Transduction
- Virus 3rd party
- Virus brings DNA from another cell
What type of horizontal gene transfer?
- Donor cell with pilus
- Fertility plasmid in donor
- Both donor and recipient alive
- Bridge forms between cells to transfer DNA
Conjugation
What type of horizontal gene transfer?
- Free donor DNA (fragment)
- Live; competent recipient cell
Transformation
What type of horizontal gene transfer?
- Donor is lysed bacterial cell; Defective bacteriophage is carrier of donor DNA; Live recipient cell of same species as donor
Transduction
What is the only type of horizontal gene transfer that is direct?
Conjugation
Some examples of products of transferred genes for conjugation include:
- Drug Resistance
- Resistance to metals
- Toxin production
- Enzymes
- Adherence molecules
- Degradation of Toxic Substances
- Uptake of Iron
Some examples of products of transferred genes for transformation include:
- Polysaccharide capsule; unlimited with cloning techniques
Some examples of products of transferred genes for transduction include:
- Toxins
- Enzymes for sugar fermentation
- Drug resistance
Conjugation
- A mode of genetic exchange in which a plasmid or other genetic material is transferred from a donor to a recipient cell via a direct connection
- Can occur in both gram-positive and gram-negative cells
An example of conjugative plasmids is:
F factor in E. coli
Describe the conjugation of the F factor:
- The F factor or the sex factor happens between a donor (marked with F+) and a recipient (marked with F-)
- There is a bridge that is created by the donor and that is by a pilus. The donor F factor is then copied and nicked, and one strand of the F factor is inserted into the recipient. And then once that F factor is copied and put into the recipient, that bridge is broken. And now that F- becomes a F+.
- For high-frequency transfer, sometimes that plasmid can be integrated into the host chromosomal DNA. So instead of having a separate plasmid, now you will have an actual integrated F-factor inside of host chromosome.
- Conjugation only happens when you have a F factor that is part of the plasmid. The plasmid is transferred into recipient.
Conjugation: Resistance (R) plasmids or factors carry what type of genes?
- Carry genes for resisting antibiotics or other drugs
- Commonly shared among bacteria through conjugation
- Can confer multiple resistance to antibiotics
- R factors can also carry genetic codes for resistance to heavy metals or for synthesizing virulence factors
Transformation
- The acceptance by a bacterial cell of small fragments of soluble DNA from the surrounding environment
- Competent: cells that are capable of accepting genetic material through transformation
- This is the uptake of free, usually chromosomal DNA, into a recipient and it’s also eventually integrated.
- Once a cell dies, it releases its contents and gets inside of the new host (recipient). Sometimes, it can get integrated into bacterial chromosomes or can be a stand alone plasmid.
How was transformation discovered?
- Worked with encapsulated and nonencapsulated Streptococcus pneumoniae and laboratory mice
- Encapsulated strains have a smooth (S) colony appearance
- Strains lacking a capsule have a rough (R) appearance
Transduction
- The process by which a bacteriophage serves as a carrier of DNA from a donor cell to a recipient cell
- Occurs naturally in a broad spectrum of bacteria
- The participating bacteria in a single transduction event must be the same species
There are two versions of transduction:
- Generalized transduction
- Specialized transduction
In generalized transduction, ...
- Random fragments of disintegrating host DNA are taken up by a phage during assembly
- Any gene from the bacterium can be transmitted
Steps of Generalized Transduction
Cell A
- A phage infects cells A (the donor cell) by injecting its DNA into it.
- While replicating its own genome and assembling new phage particles inside the bacterium, a phage particle incorporates a segment of bacterial DNA by mistake
- Cell A then lyses and releases mature phages, including the one with bacterial cell's DNA
Cell B
4. The altered phage attaches to another host cell (cell B), injecting the DNA from cell A rather than the viral nucleic acid.
5. Cell B receives this donated DNA, which recombines with its own DNA. Because the virus is defective (biologically inactive as a virus), it is unable to complete a lytic cycle. The transduced cell survives and can use this new genetic material.
What is specialized transduction?
A highly specific part of the host genome is incorporated into the virus when the prophage DNA separates from the chromosome (carrying host genes with it)
Steps of specialized Transduction
Cell A
- Phage DNA within bacterial chromosome
- Excised phage DNA contains some bacterial DNA
- New viral particles are synthesized. Some contain bacterial DNA in addition to phage DNA
- Cell A lyses and releases all new bacteriophages
Cell B
5. Infection of recipient cell transfers bacterial DNA to a new cell
6. Recombination results in two possible outcomes: either bacterial DNA or a combination of viral and bacterial DNA being incorporated into the bacterial chromosome.
What are transposable elements?
“Jumping genes” shift from one part of the genome to another
- From one chromosomal site to another
- From a chromosome to a plasmid
- From a plasmid to a chromosome
What are types of transposable elements?
- Insertion Elements - Smallest TEs consist only of two tandem repeats
- Retrotransposon - A type of TE that can transcribe DNA into RNA and then back into DNA for insertion in new location
- Other TEs contain genes that code for antibiotic resistance or toxin production
What are some general effects of transposable elements?
Scramble genetic language
Can be beneficial or adverse, depending on:
- Where the insertion occurs in a chromosome
- What kind of genes are relocated
- The type of cell involved
What are effects of transposable elements in bacteria?
- Changes in colony morphology, pigmentation, and antigenic characteristics
- Replacement of damaged DNA
- Transfer of drug resistance between bacteria
What are pathogenicity islands?
Contain multiple genes that are coordinated to create a new trait in the bacterium, making it pathogenic
Examples of PAIs include:
- Ability to scavenge iron in Yersinia pestis
- Ability to produce exotoxins Staphylococcus aureus
Mutation
- Any change to the nucleotide sequence in the genome
- The driving force of evolution
- In microorganisms, mutations become evident in altered gene expression, such as altered pigment production or development of resistance to a drug
Wild Type
- A microorganism that exhibits a natural, nonmutated characteristic
- The trait present in the highest numbers in a population
Mutant Strain
An organism that bears a mutation
What are two causes of mutations?
- Spontaneous mutation: A random change in the DNA arising from errors in replication that occur randomly
- Induced mutations: Result from exposure to known physical or chemical agents that damage DNA (known as mutagens)
Point Mutations
- Small mutations that affect only a single base on a gene
- Involve addition, deletion, or substitution of single bases
Three types of point mutations:
- Missense
- Silent
- Nonsense
Lethal Mutations
Mutations that lead to cell dysfunction or death
Neutral Mutation
Produce neither adverse nor helpful changes
Missense Mutation
Any change in the code that leads to the placement of a different amino acid
Nonsense Mutation
Changes a normal codon into a stop cod
Silent Mutation
- Alters a base but does not change the amino acid
- The redundancy of the code assures that certain amino acids will not be altered by a change in the third base of the codon
Back Mutation
Occurs when a gene that has undergone a mutation reverses to its original base composition
This type of mutation occurs when one or more bases are inserted into or deleted from a newly synthesized DNA strand
Frameshift Mutation
Frameshift Mutation
- This alters the reading frame of the mRNA
- Nearly always result in a nonfunctional protein
- Every amino acid after the mutation is different from what is coded for in the original DNA
- Insertion of bases in multiples of three does not disturb the reading frame
Photoactivation
- Light repair of damage caused by ultraviolet radiation
- Requires visible light and a light-sensitive enzyme called DNA photolyase, which can detect and attach to the damaged areas
Mismatch Repair
- A repair system can locate mismatched bases that were missed during proofreading
- The base must be replaced soon after the mismatch is made, or the repair enzymes will not recognize it
Excision Repair
- Mutations are excised by a series of enzymes that remove the incorrect bases and add the correct ones
- These bases are modified (deamination/oxidation)
What is the Ames Test?
Used to rapidly detect chemicals with carcinogenic potential
- Uses bacteria rather than experimental animals
- Allows easy observation and monitoring of gene expression and mutation rate
- Commonly uses Salmonella typhimurium because of the back mutation potential to synthesize the amino acid histidine
How is Ames test done?
- There is one culture of Salmonella that lacks histidine (-)
- Then, there are two setups. In the control plate, the Salmonella is plated on a histidine-free medium containing liver enzymes but no test agent. In the experimental/test plate, it is prepared the same way but now it contains the test agent.
- Incubate for 12 hours. Any colonies that for have back-mutated to his(+)
What is the Ames test used for?
To test whether something is a mutagen or not
Positive and Negative Effects of Mutations
Many mutations are not repaired
- Effects of mutations depend on the nature of the mutation and the strategies available to the organism
Mutations are permanent and heritable
- Passed on to the offspring of organisms and new viruses and become a long-term part of the gene pool
A small number of mutations contribute the success of the individual and the population
- Variant strains with alternate ways of expressing a trait can more readily adapt, survive, and reproduce