Instructions for Side by Side Printing
- Print the notecards
- Fold each page in half along the solid vertical line
- Cut out the notecards by cutting along each horizontal dotted line
- Optional: Glue, tape or staple the ends of each notecard together
BMD 330 Chapter 6 - Microbial Genetics
front 1 True or False: Genetics only takes place at the DNA level. | back 1 False
|
front 2 Organismal Genetics | back 2 Observes heredity of the whole organism |
front 3 Cellular Genetics | back 3 Looks at single cells |
front 4 Chromosomal Genetics | back 4 Examines the characteristics and actions of chromosomes |
front 5 Molecular Genetics | back 5 Deals with biochemistry of genes |
front 6 _______ is the sum total of genetic material of an organism. | back 6 Genome is the sum total of genetic material of an organism. |
front 7 Most of the genome exists in the form of ________ while some may appear in ______________ forms (plasmids, organelles) | back 7 Most of the genome exists in the form of chromosomes while some may appear in nonchromosomal forms (plasmids, organelles) |
front 8 ______ tiny extra pieces of DNA | back 8 Plasmids |
front 9 What two organelles have their own DNA? | back 9
|
front 10 Genomes of cells are made of ____, but viruses contain ______ or _____. | back 10 Genomes of cells are made of DNA, but viruses contain either DNA or RNA. |
front 11 The study of an organism's entire genome is known as... | back 11 Genomics |
front 12 _______ is a distinct cellular structure composed of a neatly packaged DNA molecule. | back 12 Chromosome |
front 13 How do eukaryotic and bacterial chromosome differ? | back 13
|
front 14 ____ site on the chromosome that provides basic information for a certain cell function. | back 14 Gene |
front 15 Genes fall into three categories: | back 15
|
front 16 _________ is the sum of all types of genes constituting an organism's distinctive genetic makeup. | back 16 Genotype is the sum of all types of genes constituting an organism's distinctive genetic makeup. |
front 17 _______ is the expression of the genotype that creates certain structures or functions (traits). | back 17 Phenotype is the expression of the genotype that creates certain structures or functions (traits). |
front 18 True or False: All organisms contain more genes in their phenotype than are manifested as a genotype at any given time. | back 18 False All organisms contain more genes in their genotypes than are manifested as a phenotype at any given time |
front 19 What does this statement mean?
| back 19 The phenotype can change depending on which genes are "turned on" (expressed) |
front 20 The basic unit of a DNA is.... | back 20 Nucleotide |
front 21 A nucleotide consists of: | back 21
|
front 22 What are the two types of nitrogenous bases? | back 22
|
front 23 What is an important characteristic of DNA? | back 23 DNA is arranged antiparallel
|
front 24 Two significances of the antiparallel DNA structure | back 24
|
front 25 Most DNA is replicated in a _____-_________ manner, meaning that one strand will always be the original strand and the other strand will be the newly synthesized strand. | back 25 Semi-Conservative |
front 26 Steps of the replication (simplified) | back 26
|
front 27 Helicase | back 27 Unzipping the DNA helix |
front 28 Primase | back 28 Synthesizing RNA primer |
front 29 DNA Pol III | back 29 Adding bases to the new DNA chain and proofreading the chain for mistakes |
front 30 DNA Pol I | back 30 Removing primer, closing gaps, and repairing mismatches |
front 31 Ligase | back 31 Final binding of nicks in DNA during synthesis and repair |
front 32 Topoisomerase I | back 32 Making single-stranded DNA breaks to relieve supercoiling at origin |
front 33 Topoisomerase II (DNA gyrase) and 4 | back 33 Making double-stranded DNA breaks to remove supercoiling ahead of origin and separate replicated daughter DNA molecules |
front 34 In the semi-conservative model... | back 34
|
front 35 True or False: DNA pol III can only add nucleotides in 3' to 5' direction. | back 35 False
|
front 36 Functions of DNA pol lll: | back 36
|
front 37 Replication Fork | back 37
|
front 38 Primer | back 38
|
front 39 True or False: Both eukaryotes and prokaryotes have replication forks. | back 39 True
|
front 40 The strand of new DNA that is synthesized continuously in a 5' to 3' direction | back 40 Leading Strand |
front 41 The strand of new DNA that must be synthesized in short segments (in a 5' to 3' direction) | back 41 Lagging Strand
|
front 42 What are Okazaki fragments? | back 42 Short segments of DNA synthesized in a 5' to 3' direction which are then sealing together to form the 3' to 5' strand |
front 43 DNA Replication Steps | back 43 1. During replication topoisomerases unwind the DNA helix, giving access to helicase (unzipping enzymes) to bind to the dsDNA at the origin. Helicases break the hydrogen bonds holding the two strands together, resulting in two separate strand. 2. Single-stranded binding proteins keep the strand apart. 3. DNA pol III adds nucleotides in accordance with teh template pattern. Note that RNA primase will have already added a short length of RNA. Because DNA polymerase is correctly oriented for synthesis only in the 5' to 3' direction of the new molecule strand, only one strand, called the leading strand, can be synthesized as a continuous, complete strand. The strand with the opposite orientation (3' to 5') is the lagging strand.
|
front 44 ____ begin linking fragments, and ___________ causes a double-stranded DNA break separating the two fully replicated daughter molecules. | back 44 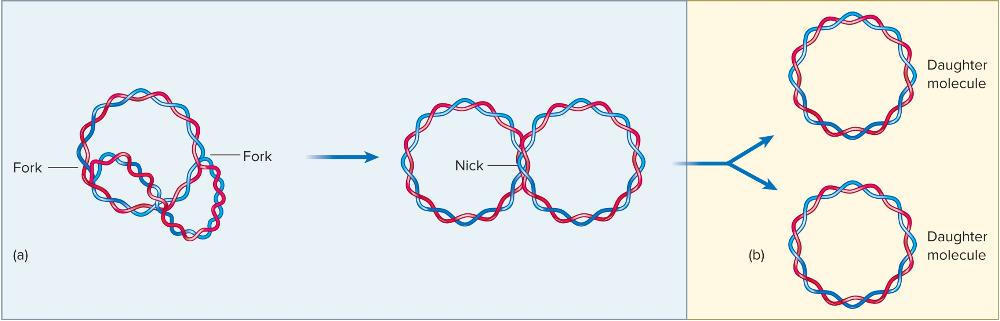 Ligases begin linking fragments, and Topoisomerase IV causes a double-stranded DNA break separating the two fully replicated daughter molecules. |
front 45 How is replication of eukaryotic DNA similar to replication of bacterial and archaeal DNA? | back 45
|
front 46 Due to the unidirectional action of DNA polymerase, the 3' end of linear DNA molecules, called ______, cannot be copied entirely | back 46 Due to the unidirectional action of DNA polymerase, the 3' end of linear DNA molecules, called telomeres, cannot be copied entirely |
front 47 _____________: DNA is used to synthesize RNA _____________: RNA used to produce proteins | back 47 Transcription: DNA is used to synthesize RNA Translation: RNA used to produce proteins |
front 48 Why is the central dogma incomplete? | back 48
|
front 49 What is the central dogma? | back 49 DNA --> RNA --> Protein |
front 50 Describe the flow of genetic information in microbes. | back 50
|
front 51 What is the connection between gene and protein? In other words, what is the connection between DNA and an organism's trait? | back 51
|
front 52 The DNA molecule is a continuous chain of base pairs, but the sequence must be interpreted in groups of ____ base pairs (a ____). Each _____ as copied into ____ codons will translate into one amino acid. | back 52 The DNA molecule is a continuous chain of base pairs, but the sequence must be interpreted in groups of three base pairs (a triplet). Each triplet as copied into mRNA codons will translate into one amino acid. |
front 53 What are the major participants in transcription and translation? | back 53
|
front 54 What is the only type of RNA that is translated? | back 54 mRNA (Messenger RNA) |
front 55 Description and Function of mRNA | back 55
|
front 56 Description and Function of tRNA (Transfer RNA) | back 56
|
front 57 Description and Function of rRNA (Ribosomal RNA) | back 57
|
front 58 Description and Function of Micro (miRNA), antisense, riboswitch, and small interfering (siRNA) | back 58
|
front 59 Description and Function of Primer | back 59
|
front 60 Description and Function of Ribozymes and Spliceosomes (snRNA) | back 60
|
front 61 How are RNAs different from DNA? | back 61
|
front 62 Messenger RNA (mRNA) | back 62
|
front 63 _____ are a series of triplet bases that hold the message of the transcribed mRNA that will be translated into an amino acid. | back 63 Codons are a series of triplet bases that hold the message of the transcribed mRNA that will be translated into an amino acid. |
front 64 tRNA are known as the _______ molecule. | back 64 Adapter |
front 65 Which type of RNA contains sequences of bases that form hydrogen bonds with complementary sections of the same tRNA strand? | back 65 Transfer RNA (tRNA) |
front 66 Where is the anticodon found? | back 66
|
front 67 What is the function of rRNA? | back 67 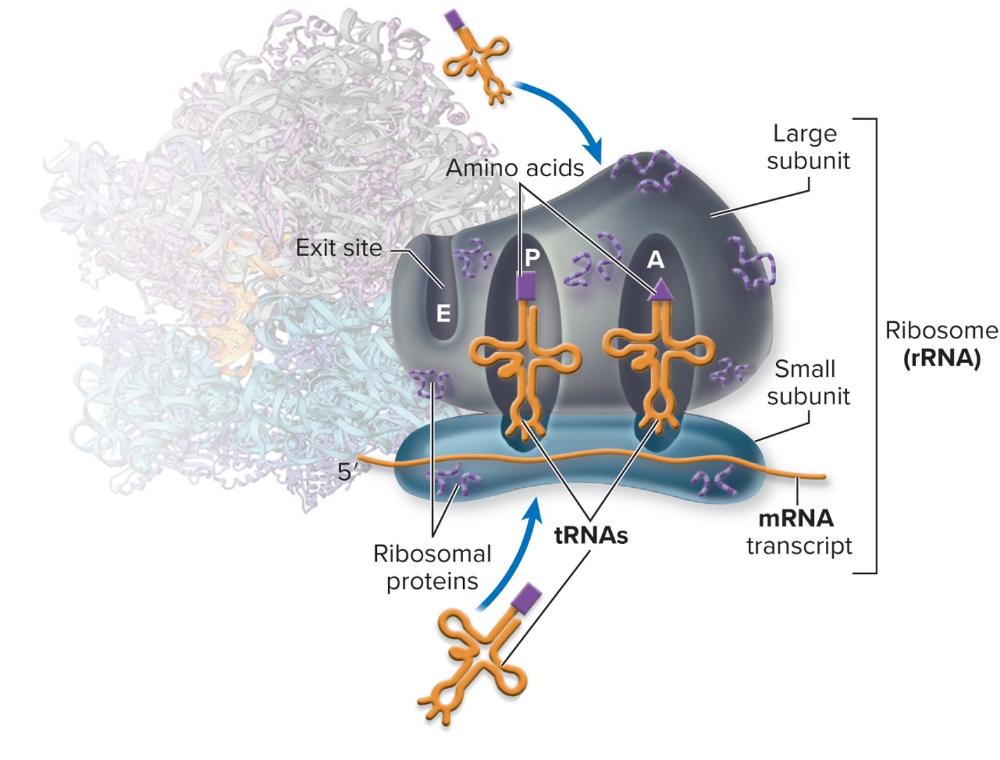
|
front 68 Three steps of transcription are: | back 68
|
front 69 When is transcription initiated? | back 69
|
front 70 What happens during the elongation step of transcription? | back 70
|
front 71 What happens during the termination step of transcription? | back 71
|
front 72 After the mRNA is made from _______, the mRNA is now ready to be _________. | back 72 After the mRNA is made from transcription, the mRNA is now ready to be translated. |
front 73 Central principle of translation: | back 73
|
front 74 In prokaryotes, the start codon is... | back 74 AUG which is f-methionine |
front 75 What are the three stop codons? | back 75 UGA, UAG, UAA |
front 76 The master genetic code is: | back 76
|
front 77 What is meant by the redundancy concept of codons? | back 77
|
front 78 What is meant by the wobble hypothesis? | back 78
|
front 79 What are the three stages of translation? | back 79
|
front 80 What are all the elements needed to synthesize a protein? | back 80
|
front 81 Start Codon | back 81
|
front 82 Stop Codon | back 82
|
front 83 Translocation | back 83
|
front 84 Can tRNA bind without the presence of ribosome? | back 84 NO |
front 85 Brief Steps of Translation | back 85 1. Entrance of tRNAs 1 and 2 2. Formation of peptide bond 3. Discharge of tRNA 1 at E site 4. First translocation 5. Formation of peptide bond 6. Discharge of tRNA 2; second translocation; enter tRNA 4 7. Formation of peptide bond |
front 86 A little more in depth of translation: | back 86
|
front 87 Explain the entirety of translation | back 87 1. The mRNA molecule leaves the DNA transcription site and is transported to ribosomes in the cytoplasm. Ribosomal subunits come together and form sites to hold the mRNA and tRNAs. The ribosome begins to scan the mRNA by moving in the 5' to 3' direction along the mRNA. The first codon it encounters called the start codon. With the mRNA message in place on the assembled ribosome, the next step in translation involves entrance of tRNAs with their amino acids. The pool of cytoplasm contains a complete array of tRNAs, previously charged by having the correct amino acid attached. The step in which the complementary tRNA meets the mRNA code is guided by the two sites on the large subunit of the ribosome called the P site and the A site. The ribosome also has an exit or E site where used tRNAs are released. 2. Rules of pairing dictate that the anticodon of tRNA must be complementary to the mRNA codon. The ribosome shifts its "reading frame" to the right along the mRNA from one codon to the next. This brings the next codon into place on the ribosome and makes a place for the next tRNA to enter the A position. A peptide bond is formed between the amino acids on the adjacent tRNAs, and the polypeptide grows. Elongation beings with the filling of the A site by a second tRNA. The identify of this tRNA and its amino acid is dictated by the second mRNA codon. 3. The entry of tRNA into the A site brings two adjacent tRNAs in favorable proximity for a peptide bond to form between the amino acids they carry. For the next step to proceed, some room must be made on the ribosome, and the next codon in sequence must be brought into position for reading. This process is accomplished by translocation, the enzyme-directed shifting of the ribosome to the right along the mRNA strand, which causes the blank tRNA 1 to be discharged from the ribosome at the E site. 4. Dipeptide moves to P site. Site A is empty. tRNA that has been released is now free to drift off into the cytoplasm and become recharged with an amino acid for later additions to this or another protein. The stage is now set for the insertion of tRNA 3 at site A as directed by the third mRNA codon. This insertion is followed once again by peptide bond formation between the dipeptide and amino acid 3 (making a tripeptide), splitting of the peptide from tRNA 2, and translocation. 5. Releases tRNA 2, shifts mRNA to next position, moves tRNA 3 to position P, and opens position A for next tRNA. 6. From this point on, peptide elongation proceeds. 7. The termination is brought by presence of at least one special codon occurring just after the codon for the last amino acid. When this codon is reached, a special enzyme breaks the bond between the final tRNA and the finished polypeptide chain, releasing it from the ribosome. |
front 88 What happens after protein synthesis? | back 88 Posttransitional modifications
|
front 89 Why is protein assembly line in bacteria faster? | back 89
|
front 90 COVID-19 viruses have... | back 90
|
front 91 How does the COVID-19 mRNA vaccine work? | back 91 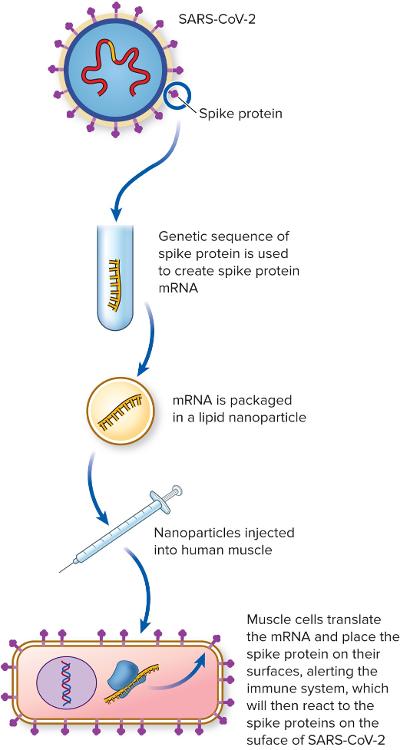
|
front 92 What are differences between prokaryotic and eukaryotic transcription and translation? | back 92
|
front 93 ____ are the coding regions whereas the _____ are intervening sequences of bases that do not code for protein. | back 93 Exons are the coding regions whereas the introns are intervening sequences of bases that do not code for protein. |
front 94 What recognizes the exon-intron junctions and enzymatically cuts them? | back 94 Spliceosome
|
front 95 Operons | back 95
|
front 96 Inducible Operons | back 96
|
front 97 Repressible Operons | back 97
|
front 98 What type of operon contains genes that make proteins that help metabolize nutrients? | back 98 Inducible Operons |
front 99 What type of operon contain genes that synthesize amino acids | back 99 Repressible |
front 100 Three features of the lac operon: | back 100
|
front 101 Regulator | back 101 Composed of the gene that codes for the repressor, a protein capable of repressing the operon |
front 102 Control Locus | back 102
|
front 103 Structural Locus | back 103 Made up of three genes, each coding for a different enzyme needed to catabolize lactose |
front 104 What is the main purpose of the lactose operon? | back 104
|
front 105 The repressor protein for the lactose operon is... | back 105 Allosteric |
front 106 What does allosteric protein do in a lactose operon? | back 106 It is the repressor protein:
|
front 107 Describe the lac operon in bacteria | back 107 1. This operon is normally in an "off" mode and does not initiate transcription when the appropriate substrate is absent. 2. If lactose is added to the cell's environment, it triggers events that turn the operon on. 3. The structural genes are transcribed in a single unbroken transcript coding for all three enzymes. (During translation, however, each protein is synthesized separately). |
front 108 The lac operon is usually... inducible. | back 108
|
front 109 Repressible operon | back 109
|
front 110 Repressible Operon: Control of a Gene Through Excess Nutrients | back 110
|
front 111 What two ways do eukaryote cells regulate genes? | back 111
|
front 112 Transcription Factors | back 112
|
front 113 DNA "knot" | back 113
|
front 114 Phase Variation | back 114 The result of bacteria turning on or off a complement of genes that leads to phenotypic changes
|
front 115 What are the drugs the interfere with ribosome? | back 115
|
front 116 What are the drugs that inhibit protein synthesis? | back 116
|
front 117 An event in which one bacterium donates DNA to another bacterium. | back 117 Recombination |
front 118 What is the result of recombination? | back 118 The end result is a strain different from both the donor and the recipient strain |
front 119 What is the extrachromosomal DNA adept at moving between cells called? | back 119 Plasmids |
front 120 Any organism that contains and expresses genes that originated in another organism is known as... | back 120 Recombinant |
front 121 What is horizontal gene transfer? | back 121 Any transfer of DNA that results in organisms acquiring new genes that did not come directly from parent organisms |
front 122 Horizontal gene transfer does not include chromosomes. It is about ______. | back 122 Plasmids |
front 123 Can eukaryotic organisms also engage in horizontal gene transfer? | back 123 Eukaryotic organisms also engage in horizontal gene transfer, often aided by microbes such as viruses |
front 124 What are the three types of horizontal gene transfer? | back 124
|
front 125 Small, circular pieces of DNA that can replication independently of the bacterial chromosome are... | back 125 Plasmids
|
front 126 Conjugation | back 126
|
front 127 Transformation | back 127
|
front 128 Transduction | back 128
|
front 129 What type of horizontal gene transfer?
| back 129 Conjugation |
front 130 What type of horizontal gene transfer?
| back 130 Transformation |
front 131 What type of horizontal gene transfer?
| back 131 Transduction |
front 132 What is the only type of horizontal gene transfer that is direct? | back 132 Conjugation |
front 133 Some examples of products of transferred genes for conjugation include: | back 133
|
front 134 Some examples of products of transferred genes for transformation include: | back 134
|
front 135 Some examples of products of transferred genes for transduction include: | back 135
|
front 136 Conjugation | back 136
|
front 137 An example of conjugative plasmids is: | back 137 F factor in E. coli |
front 138 Describe the conjugation of the F factor: | back 138
|
front 139 Conjugation: Resistance (R) plasmids or factors carry what type of genes? | back 139
|
front 140 Transformation | back 140
|
front 141 How was transformation discovered? | back 141
|
front 142 Transduction | back 142
|
front 143 There are two versions of transduction: | back 143
|
front 144 In generalized transduction, ... | back 144
|
front 145 Steps of Generalized Transduction | back 145 Cell A
Cell B 4. The altered phage attaches to another host cell (cell B), injecting the DNA from cell A rather than the viral nucleic acid. 5. Cell B receives this donated DNA, which recombines with its own DNA. Because the virus is defective (biologically inactive as a virus), it is unable to complete a lytic cycle. The transduced cell survives and can use this new genetic material. |
front 146 What is specialized transduction? | back 146 A highly specific part of the host genome is incorporated into the virus when the prophage DNA separates from the chromosome (carrying host genes with it) |
front 147 Steps of specialized Transduction | back 147 Cell A
Cell B 5. Infection of recipient cell transfers bacterial DNA to a new cell 6. Recombination results in two possible outcomes: either bacterial DNA or a combination of viral and bacterial DNA being incorporated into the bacterial chromosome. |
front 148 What are transposable elements? | back 148 “Jumping genes” shift from one part of the genome to another
|
front 149 What are types of transposable elements? | back 149
|
front 150 What are some general effects of transposable elements? | back 150 Scramble genetic language Can be beneficial or adverse, depending on:
|
front 151 What are effects of transposable elements in bacteria? | back 151
|
front 152 What are pathogenicity islands? | back 152 Contain multiple genes that are coordinated to create a new trait in the bacterium, making it pathogenic |
front 153 Examples of PAIs include: | back 153
|
front 154 Mutation | back 154
|
front 155 Wild Type | back 155
|
front 156 Mutant Strain | back 156 An organism that bears a mutation |
front 157 What are two causes of mutations? | back 157
|
front 158 Point Mutations | back 158
|
front 159 Three types of point mutations: | back 159
|
front 160 Lethal Mutations | back 160 Mutations that lead to cell dysfunction or death |
front 161 Neutral Mutation | back 161 Produce neither adverse nor helpful changes |
front 162 Missense Mutation | back 162 Any change in the code that leads to the placement of a different amino acid |
front 163 Nonsense Mutation | back 163 Changes a normal codon into a stop cod |
front 164 Silent Mutation | back 164
|
front 165 Back Mutation | back 165 Occurs when a gene that has undergone a mutation reverses to its original base composition |
front 166 This type of mutation occurs when one or more bases are inserted into or deleted from a newly synthesized DNA strand | back 166 Frameshift Mutation |
front 167 Frameshift Mutation | back 167
|
front 168 Photoactivation | back 168
|
front 169 Mismatch Repair | back 169
|
front 170 Excision Repair | back 170
|
front 171 What is the Ames Test? | back 171 Used to rapidly detect chemicals with carcinogenic potential
|
front 172 How is Ames test done? | back 172
|
front 173 What is the Ames test used for? | back 173 To test whether something is a mutagen or not |
front 174 Positive and Negative Effects of Mutations | back 174 Many mutations are not repaired
Mutations are permanent and heritable
A small number of mutations contribute the success of the individual and the population
|