1) What is metagenomics?
A) genomics as applied to a species
that most typifies the average phenotype of its genus B) the
sequencing of one or two representative genes from several
species
C) the sequencing of only the most highly conserved
genes in a lineage
D) sequencing DNA from a group of species
from the same ecosystem
Answer: D
2) An early step in shotgun sequencing is to _____.
A) break
genomic DNA at random sites
B) map the position of cloned DNA
fragments
C) randomly select DNA primers and hybridize these to
random positions of chromosomes in preparation for sequencing
Answer: A
3) Using modern techniques of sequencing by synthesis and the shotgun
approach, sequences are assembled into chromosomes by ______.
A)
placing them on previously generated genetic maps.
B) cloning
them into plasmid vectors.
C) computer analysis looking for sequence overlaps.
D) cloning
them into plasmid vectors, placing them on previously generated
genetic maps, followed by computer analysis looking for sequence overlaps.
Answer: C
4) Proteomics is defined as the _____.
A) linkage of each gene
to a particular protein
B) study of the full protein set encoded
by a genome
C) totality of the functional possibilities of a
single protein
D) study of how amino acids are ordered in a protein
Answer: B
5) Bioinformatics can be used to scan for short sequences that
specify known mRNAs, called _____.
A) expressed sequence
tags
B) multigene families
C) proteomes
D) short tandem repeats
Answer: A
6) What is gene annotation in bioinformatics?
A) finding
transcriptional start and stop sites, RNA splice sites, and ESTs in
DNA sequences B) assigning names to newly discovered genes
C)
describing the functions of noncoding regions of the genome
D)
matching the corresponding phenotypes of different species
Answer: A
7) Bioinformatics includes _____.
I. using computer programs to
align DNA sequences
II. creating recombinant DNA from separate
species
III. developing computer-based tools for genome
analysis
IV. using mathematical tools to make sense of
biological systems
A) I and II
B) II and III
C) II and IV
D) I, III,
and IV
Answer: D
8) After finding a new medicinal plant, a pharmaceutical company
decides to determine if the plant has genes similar to those of other
known medicinal plants. To do this, the company annotates the genome
of the new plant to _____.
A) determine what proteins are produced
B) determine what mRNA transcripts are produced
C) identify
genes and determine their functions
D) identify the location of
mRNA within the plant cells
Answer: C
9) If the sequence of a cDNA has matches with DNA sequences in the
genome, then this genomic DNA is likely to _____.
A) code for a
protein
B) code for an rRNA
C) be part of an intron
D) be a regulatory sequence
Answer: A
10) In what sense are studies by nineteenth-century naturalists and
those by early twenty-first- century genomic biologists similar?
A) Both focused on cellular evolution.
B) Both worked to
understand how genes evolved.
C) Both took a reductionist approach by studying only one small part of a complex system.
D) Both focused on observing and describing what exists in their realms of investigation.
Answer: D
11) Which of the following techniques would be most appropriate to
test the hypothesis that humans and chimps differ in the expression of
a large set of shared genes?
A) DNA microarray analysis
B)
polymerase chain reactin (PCR)
C) DNA sequencing
D) protein-protein interaction assays
Answer: A
12) A DNA microarray is a tool that owes its existence to earlier
genomics investigations. What essential contribution of genomics makes
microarrays possible?
A) recently improved RNA sequencing
technologies
B) continuously improving methods of gene cloning
C) more efficient techniques for cDNA synthesis
D) knowledge
of which DNA sequences to synthesize for the array
Answer: D
13) What can proteomics reveal that genomics cannot?
A) the
number of genes characteristic of a species
B) the patterns of
alternative splicing
C) the set of proteins present within a
cell or tissue type
D) the movement of transposable elements
within the genome
Answer: C
14) A sequence database such as GenBank could be used to do all of
the following EXCEPT A) Compare cow and human insulin protein
sequences.
B) Construct a tree to determine the evolutionary
relationships between various bird species.
C) Search for genes in a fruit fly that are similar to a human gene.
D) Compare patterns of gene expression in cancerous and non-cancerous cells.
Answer: D
15) Current analysis indicates that less than 2% of the human genome codes for proteins. Based on the systems approach employed by the ENCODE project, what percentage of the genome is estimated to contain functional elements (includes functional RNAs and regulatory sequences)?
A) Less than 2%
B) About 50%
C) At least 80%
D) 100%
Answer: C
16) Which of the following is a representation of gene density?
A) Humans have 2900 Mb per genome.
B) C. elegans has ~20,000
genes.
C) Humans have ~20,000 protein-encoding genes in 2900 Mb.
D) Fritillaria has a genome 40 times the size of a human.
Answer: C
17) Why might the cricket genome have eleven times as many base pairs
as that of Drosophila melanogaster?
A) Crickets have higher gene
density.
B) Drosophila are more complex organisms.
C) Crickets must have more noncoding DNA.
D) Crickets must make many more proteins.
Answer: C
18) The comparison between the number of human genes and those of
other animal species has led to many conclusions, including that
_____.
A) the density of the human genome is far higher than in
most other animals
B) the number of proteins expressed by the
human genome is far more than the number of its genes
C) most human DNA consists of genes for protein, tRNA, rRNA, and
miRNA
D) the genomes of most other organisms are significantly
smaller than the human genome
Answer: B
19) It is more difficult to identify eukaryotic genes than
prokaryotic genes because in eukaryotes _____.
A) the proteins
are larger than in prokaryotes
B) the coding portions of genes
are shorter than in prokaryotes
C) there are no start codons
D) there are introns
Answer: D
20) If alternative splicing did NOT occur then _____.
A) the
human genome would likely contain many more genes
B) the E. coli
genome would contain many fewer genes
C) there would be little
correlation between the complexity of organisms and genome size
D) there would be many fewer genes devoted to metabolism in Arabidopsis and yeast
Answer: A
21) A multigene family is composed of _____.
A) multiple genes
whose products must be coordinately expressed
B) genes whose
sequences are very similar and that probably arose by duplication
C) a gene whose exons can be spliced in a number of different
ways
D) a highly conserved gene found in a number of different species
Answer: B
22) Retrotransposons _____.
A) use an RNA molecule as an
intermediate in transposition
B) are found only in animal
cells
C) generally move by a cut-and-paste mechanism
D)
contribute a significant portion of the genetic variability seen
within a population of gametes
Answer: A
23) In humans, the embryonic and fetal forms of hemoglobin have a
higher affinity for oxygen than that of adults. This is due to
_____.
A) nonidentical genes that produce different versions of
globins during development.
B) pseudogenes, which interfere with
gene expression in adults.
C) the attachment of methyl groups to cytosine following birth,
which changes the type of hemoglobin produced.
D) histone
proteins changing shape during embryonic development.
Answer: A
24) Sequencing eukaryotic genomes is more difficult than sequencing
genomes of bacteria or archaea because of the _____.
A) large
size of eukaryotic proteins
B) hard-to-find proteins
C) high proportion of G-C base pairs in eukaryotic DNA
D)
large size of eukaryotic genomes and the large amount of eukaryotic
repetitive DNA
Answer: D
25) Because they both produce a reverse transcriptase, long
interspersed nuclear elements (LINEs) as transposable elements may be
related to _____.
A) plasmids
B) retroviruses
C) poliovirus
D) parasitic bacteria
Answer: B
26) Although transposable elements and short tandem repeats (STRs)
are repetitive DNAs, they differ in that _____.
A) STRs occur
within exons; transposable elements occur within introns
B) STRs
occur within introns; transposable elements occur within exons
C) the repeated unit in STRs is clustered one after another;
transposable element repeats are scattered throughout the genome
D) the repeated unit in STRs is much larger than the repeated unit of
transposable elements
Answer: C
27) Which of the following can be duplicated in a genome?
A) only DNA sequences
B) only entire sets of chromosomes
C) only entire chromosomes
D) DNA sequences, chromosomes, or sets of chromosomes
Answer: D
28) Unequal crossing over during prophase I can result in one sister chromosome with a deletion and another with a duplication. A mutated form of hemoglobin, so-called hemoglobin Lepore, exists in the human population. Hemoglobin Lepore has a deleted series of amino acids. If this mutated form was caused by unequal crossing over, what would be an expected consequence?
A) There should also be persons whose hemoglobin contains two copies of the series of amino acids that is deleted in hemoglobin Lepore.
B) Each of the genes in the hemoglobin gene family must show the same deletion.
C) The deleted gene must have undergone exon shuffling.
D) The
deleted region must be located in a different area of the individual's genome.
Answer: A
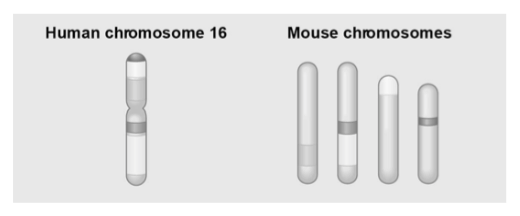
The figure above shows a diagram of blocks of genes on human chromosome 16 and the locations of blocks of similar genes on four chromosomes of the mouse.
29) The movement of these blocks suggests that _____.
A)
during evolutionary time, these sequences have separated and have
returned to their original positions
B) DNA sequences within
these blocks have become increasingly divergent
C) sequences
represented have duplicated at least three times
D) chromosomal
translocations have moved blocks of sequences to other chromosomes
Answer: D
30) Humans have twenty-three pairs of chromosomes and chimps have
twenty-four pairs of chromosomes. What is the most likely explanation
for these differences in human and chimp genomes?
A) The common
ancestor of humans and chimps had twenty-four pairs of chromosomes.
After the two groups evolved, two human chromosomes fused end to end
B) In the evolution of chimps, new adaptations resulted from
additional chromosomal material. C) At some point in evolution, human
and chimp ancestors reproduced with each other.
D) Errors in
mitosis resulted in an additional pair of chromosomes in chimps.
Answer: A
31) Exon shuffling occurs during _____.
A) splicing of DNA
B) DNA replication
C) meiotic recombination
D) translation
Answer: C
32) When gene duplication occurs to its ultimate extent by doubling
all genes in a genome, what has occurred?
A) pseudogene
creation
B) creation of a gene cluster
C) creation of a polyploid
D) creation of a diploid
Answer: C
33) Mutations that occur in one member of a gene pair that arose from
gene duplication may create _____.
A) a pseudogene
B) a
gene with a new function
C) a gene family with two distinct but related members
D) a
pseudogene, a gene with a new function, and a gene family with two
distinct but related members
Answer: D
34) Based on the data in the Amino Acid Identity Table, which two
members of the human globin gene family are the most divergent?
A) a1 and ß
B) ? and ß
C) a1 and a2
D) a1 and G?
Answer: B
35) Fragments of DNA have been extracted from the remnants of extinct
woolly mammoths, amplified, and sequenced. These can now be used to
_____.
A) introduce certain mammoth traits into relatives, such
as elephants
B) clone live woolly mammoths
C) appreciate the reasons why mammoths went extinct
D) better
understand the evolutionary relationships among members of related taxa
Answer: D
36) Homeotic genes contain a homeobox sequence that is highly
conserved among very diverse species. The homeobox is the code for
that domain of a protein that binds to DNA in a regulatory
developmental process. Therefore, you would expect that _____.
A) homeotic genes are selectively expressed as an organism develops
B) homeoboxes cannot be expressed in nonhomeotic genes
C) homeotic genes in apes and humans are very different
D) all organisms must have homeotic genes
Answer: A
37) A recent study compared the Homo sapiens genome with that of
Neanderthals. The results of the study indicated that there was a
mixing of the two genomes at some period in evolutionary history.
Additional data consistent with this hypothesis could be the discovery
of _____.
A) some Neanderthal sequences not found in living humans
B) a few modern H. sapiens with some Neanderthal sequences
C)
duplications of several Neanderthal genes on a Neanderthal
chromosome
D) some Neanderthal chromosomes that are shorter than
their counterparts in living humans
Answer: B
38) Several of the different globin genes are expressed in humans,
but at different times in development. What mechanism could allow for
this?
A) exon shuffling
B) pseudogene activation
C) differential translation of mRNAs
D) differential gene
regulation over time
Answer: D
39) Biologists now routinely test for homology between genes in
different species. If genes are determined to be homologous, it means
that they are related _____.
A) by descent from a common
ancestor
B) because of convergent evolution
C) by chance mutations
D) in function but not structure
Answer: A
40) A current view of how the human and chimpanzee can share most of
their nucleotide sequence yet exhibit significant phenotypic
differences is that many of the most important sequence differences
alter _____.
A) structural genes
B) the number of repeated sequences
C) regulatory sequences
D) environmental factors
Answer: C
41) Studies in knockout mice have demonstrated an important role of the FOXP2 transcription factor in the development of vocalizations. Recent sequence comparisons of the FOXP2 gene in Neanderthals and modern humans show that while the DNA sequence may be different, the protein sequence it codes for is identical. What might logically be inferred from this information?
A) There was a problem with the experiment because different DNA
sequences cannot result in the same protein sequence.
B) The
differences in DNA sequence support the hypothesis that Neanderthals
were primitive beings that could only grunt.
C) Human and Neanderthal vocalizations may have been more similar than previously thought. D) The experiments in mice demonstrating the function of the FOXP2 gene are not relevant to humans and Neanderthals because they are not primates.
Answer: C
42) Comparisons of DNA sequences within the human species have
revealed many variations. Which of the following variations involves
duplication of relatively long stretches of chromosomes, often
including the duplication of protein-coding genes?
A) CNVs
B) SNPs
C) STRs
D) Transposable elements
Answer: A
43) A microarray is a tool used in genetic research to determine the mRNAs being produced in a particular tissue, and their relative level of expression. Known genes can therefore be assayed for their expression in different situations. One use of the technology is in cancer diagnosis and treatment. If a known gene functions as a tumor suppressor, predict which of the following pieces of evidence would be most useful in diagnosis of a cancer due to a mutation in this tumor- suppressor gene.
A) The tissue sample shows a high level of gene expression relative
to a control (noncancerous) sample.
B) The tissue sample
responds to treatment with a mitosis-promoting compound.
C) The
mRNAs for the targeted tumor suppressor sequence are not being produced.
D) The mRNAs for cyclins and kinases show unusually high levels of expression.
Answer: C