What components are found in both prokaryotic and eukaryotic cells? What are the functions of these components?
- Plasma membrane - encloses the interior of the cell, regulates flow
- Cytoplasm - thick solution that fills cell and is mainly composed of water, salts, and proteins
- Ribosomes
Prokaryotic cells: unicellular
- Nucleoid: location and structural organization of the circular chromosome
- Plasmids: small DNA molecules that that contain genes but are physically independent of the cellular chromosome
- Ribosomes: complex structures composed of large and small subunits, each of which contains RNA and protein molecules
- Photosynthetic membranes that contain enzymes and pigment
molecules required for
these reactions to occur and develop as infoldings of the plasma membrane - Cytoskeleton: Protein filaments that help maintain cell shape
- Cytoplasm: all the contents of the cell excluding the nucleus
- Cell wall: tough, fibrous layer that surrounds the plasma membrane and prevents the cell from lysing
- Many bacteria have another layer outside of the cell wall that consists of lipids with polysaccharides
- Flagella: assembled from proteins and propels cells through water
- Fimbriae: needlelike projections that extend from the plasma membrane of some bacteria and promote attachment to other cells or surfaces
Eukaryotic cells: multicellular
- Cytosol: the fluid portion between the plasma membrane and organelles
- Nucleus: contains chromosomes and functions as an administrative center for information storage and processing
- Enclosed by a double membrane called the nuclear envelope which is studded with pore like openings and the inside surface is linked to fibrous proteins that form lattice like sheets called nuclear lamina
- Nuclear lamina stiffens the structure and maintains shape
- Nucleolus: active site where RNA molecules found in ribosomes are manufactured and the large and small ribosomal units are assembled
- Ribosomes: complex macromolecular machines that manufacture proteins , they are not classified as organelles because they are not surrounded by membranes
- Endoplasmic reticulum: the portions of the nuclear envelope extend to the cytoplasm to form an extensive membrane enclosed factory
- Rough ER: named for the ribosomes that attach to the membrane and synthesize proteins that will be inserted into the plasma membrane, secreted to the cell exterior and shipped to an organelle
- The lumen is the interior of the sac like component of the rough ER and where the proteins move into after being manufactured
- Smooth ER: contains enzymes that catalyze reactions involving lipids, manufacturing site for phospholipids
- Golgi Apparatus: the products of the ER pass through the golgi apparatus before they reach their final destination and consist of discrete flattened membranous sacs called cisternae which are stacked upon each other, it has distinct polarity
- Cis surface is closest to the nucleus, receives products from the rough ER
- Trans surface is oriented toward the plasma membrane, ships products out to other organelles or cell surface
- Within the cisternae the rough ER’s products are processed and packaged
- Lysosomes: function as recycling centers, contain enzymes specialized for hydrolyzing different types of macromolecules and the products leave via transport proteins in the organelle’s membrane
- Vacuoles: Are in place instead of lysosomes in plants, fungi and other groups, storage depots
- Peroxisomes: have a single membrane and originates as buds from the ER, centers for reduction oxidation reactions
- Mitochondria: produces ATP
- Cytoskeleton: gives cell its shape and is involved in moving the cell as well as materials within the cell
- Cell wall: In fungi, algea and plants, provides structural support and prevents cell from lysing
What are the advantages of compartmentalization in organelles?
- CLIR - Clumping, Localization, Incompatibility, Reducing
- C- Clumping of certain enzymes, after one is created can be used again (efficiency)
- L - Localize certain substrates, for certain chemicals, in certain areas (specificity)
- I - Incompatible substances that are needed for 1 organelle, but harmful to others can be internalized/separated/concentrated in different areas
- R- Reducing amount of cytosol
What is the “diffusion problem” and how is it solved in eukaryotic cells?
- Problem: As the cell increases in diameter, its volume increases more than its surface area therefore the rate of exchange is decreased, diffusion only allows for rapid movement across short distances
- Solved: because the eukaryotic cells are highly compartmentalized the cytosol is only a fraction of total cell volume reducing effect of total cell surface-to-volume ratio
What components are found only in eukaryotic cells? What are the functions of these components?
- Mitochondria - produces ATP
- Nucleus - holds DNA
- Golgi Apparatus - synthesizes RNA
- lysosomes- recycling bin
- membrane bound organelles
- Microtubules
- ER - rough + smooth
- peroxisome- centers for reduction oxidation reactions
How does the nuclear envelope differ from the plasma membrane? How do molecules move between the cytoplasm and the nucleus?
- Nuclear envelope: is supported by fibrous proteins that form a lattice-like sheet called the nuclear lamina which stiffens and shapes the structure
- Plasma membrane:
- Through the nuclear pore complex mediates the selective exchange of components including RNA, ribosomal proteins, signaling molecules, and lipids between the nucleus and cytoplasm
- Through nuclear pores, nuclear localization signal (NLS)
Describe the experiment that suggested that a particular sequence of amino acids served as a kind of “zip code” to direct large proteins, such as nucleoplasmin, into the nucleus? Is the uptake of nucleoplasmin tails into the nucleus active or passive transport? How can you tell?
- Labeled nucleoplasmin (strictly found in nucleus) with a radioactive atom then injected it into the cytoplasm of the cells. It quickly then went to the nucleus.
- Then researched used enzymes to separate the core section and tails, labeled each w/ radioactive atom, then injected them into cytoplasm of diff cells
- The tail ends entered the nucleus quickly but the core could not enter
- hypothesis: Nucleoplasmin contains a discrete “send to nucleus “ signal that resides in either the tale or core region
- null hypothesis: nucleoplasmin doesn’t require a signal to enter the nucleus, or the entire protein serves as a signal
- prediction: labeled tail region or labeled core region will be found in the nucleus
- Null hypothesis prediction: Either both (no protein signal) or neither (whole protein signal)
- conclusion: send to nucleus signal is in the tail region of the nucleoplasmin
- Active because it goes against its concentration gradient.
Are signal sequences unique to nuclear proteins? If not, where else can proteins be targeted? Are the signal sequences on proteins targeted to these different compartments the same?
No, other proteins have signal NLS extracellular proteins
- Phosphorylated sugars serves as a zip codes for lysosome bound proteins
What are the main components of the endomembrane system?
- Golgi Apparatus - cis surface (closest to nucleus) receives products from rough ER. Trans side (oriented towards plasma membrane) ships products out to other organelles or cell surface + beyond. in b/t within cisternae products are processed + packaged for delivery. Ex: Acid hydrolysis processed in Golgi them shipped to lysosome
- Lysosomes - Recycling centers that hydrolyze different macromolecules and send to cytosol for new energy
- Endoplasmic reticulum - Acid hydrolysis are synthesized here processed in the golgi and then shipped to the
- Producing, processing, and transporting proteins and lipids in eukaryotic cells
What is the secretory pathway hypothesis? How was the secretory pathway hypothesis experimentally confirmed?

Protein enters RER, protein exits RER in a vesicle, protein enters golgi apparatus, protein exits golgi apparatus, protein is secreted from cell in a vesicle
What is meant by a “pulse-chase” experiment?
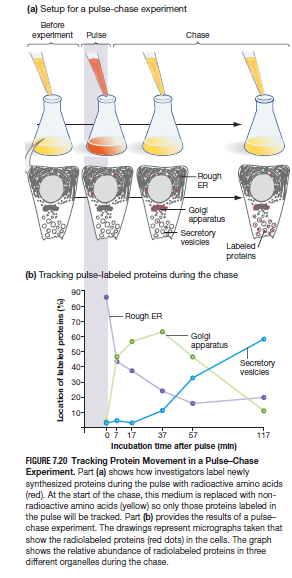
- The Pulse-Chase Method: originated from observation by George Palade of large amounts of Golgi apparatus and ER found in cells that secrete digestive enzymes, hormones, or etc. by looking through electron micrographs. Lead to hypothesis stating that cells partake in a “secretory pathway” that starts in the ER and ends in the extracellular membrane.
- Pulse - expose cell to high concentration of amino acids for a short period of time (Ex. radiolabeled amino acid)
- Chase - time following the end of the pulse. the pulse ends by washing away the modified amino acid and replacing it with the normal version of the same molecule. If the chase consists of unlabeled amino acid, then the proteins synthesized will not be radiolabeled.
- Intention - mark a particular group during an interval and follow their changes (if any) overtime specifically with pancreatic cells growing in culture which are usually packed with ER and golgi and are used in small intestine to secrete digestive enzymes
- Results - Showed that proteins are trafficked through the secretory pathway from RER->Glolgi ->extracellular in about two hours
Where in the cell are proteins synthesized? What is the “signal hypothesis,” and how was the signal hypothesis experimentally confirmed for proteins that enter the lumen of the ER? How are proteins that are produced in the ER transported to the Golgi?
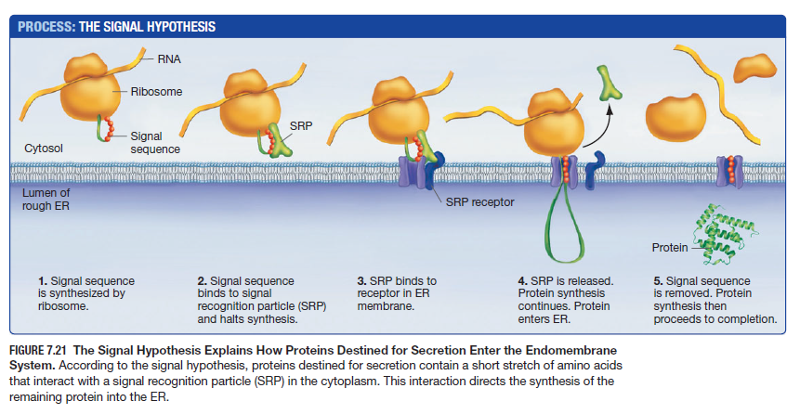
Proteins are synthesized in ribosomes free in the cytosol -> eventually these ribosomes become attached to the rough ER
Signal Hypothesis
- First few amino acids in the growing polypeptide chain act as a signal that marks the ribosome for transport to the ER membrane
- Realized that when proteins that are normally synthesized in the rough ER are manufactured by free floating ribosomes instead they are on average 20 more amino acids longer
- Prediction: Proteins bound for the endomembrane system have a molecular zipcode analogous to nuclear localization signal (NLS)
- Results: Extra amino acids on free floating ribosomes are the “send to ER” signal called ER signal sequence - will move proteins to ER lumen
Proteins are transported from the ER to the Golgi by vesicles that bud off from the rough ER and carried to the golgi apparatus with only the appropriate cargo and then fuse with the membrane of the cis face of the golgi (side nearest rough ER)
What is “receptor-mediated endocytosis?” Draw the pathway that moves molecules from the extracellular space to the lysosome.
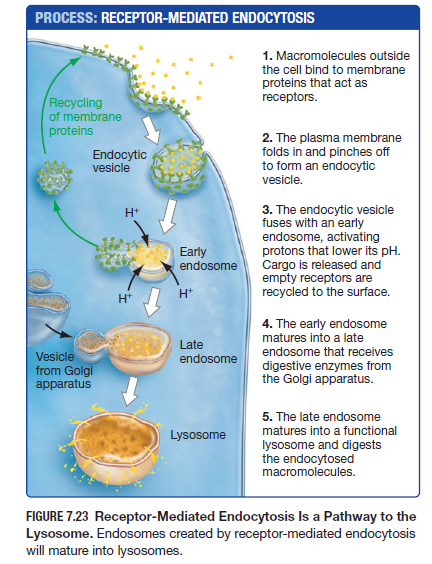
Receptor-mediated endocytosis is the most specific type of endocytosis. The main difference between receptor-mediated endocytosis and phagocytosis is that rather than the cell membrane’s specific protein receptors attaching to the receptors of the bacterial cell like in phagocytosis, in receptor-mediated endocytosis the actual molecule that is being ingested binds directly to the receptors of the cell membrane (ex. macromolecules like sugars and hormones) pinching off to form a vesicle and fuses with an early endosome to lower it’s pH and release it’s cargo before being digested and recycling its membrane proteins for another cycle of receptor-mediated endocytosis.