1) In his transformation experiments, what did Griffith
observe?
A) Mixing a heat-killed pathogenic strain of bacteria
with a living nonpathogenic strain can convert some of the living
cells into the pathogenic form.
B) Mixing a heat-killed
nonpathogenic strain of bacteria with a living pathogenic strain makes
the pathogenic strain nonpathogenic.
C) Infecting mice with
nonpathogenic strains of bacteria makes them resistant to pathogenic
strains.
D) Mice infected with a pathogenic strain of bacteria
can spread the infection to other mice.
A
2) How do we describe transformation in bacteria?
A) the
creation of a strand of DNA from an RNA molecule
B) the creation of a strand of RNA from a DNA molecule
C) the infection of cells by a phage DNA molecule
D) assimilation of external DNA into a cell
D
4) In trying to determine whether DNA or protein is the genetic
material, Hershey and Chase made use of which of the following
facts?
A) DNA contains sulfur, whereas protein does not.
B)
DNA contains phosphorus, whereas protein does not.
C) DNA contains nitrogen, whereas protein does not.
D)
DNA contains purines, whereas protein includes pyrimidines.
B
5) Which of the following investigators was (were) responsible for
the following discovery?
In DNA from any species, the amount of
adenine equals the amount of thymine, and the amount of guanine equals
the amount of cytosine.
A) Alfred Hershey and Martha
Chase
B) Oswald Avery, Maclyn McCarty, and Colin MacLeod
C)
Erwin Chargaff
D) Matthew Meselson and Franklin Stahl
C
7) It became apparent to Watson and Crick after completion of their model that the DNA molecule could carry a vast amount of hereditary information in which of the following?
A) sequence of bases
B) phosphate-sugar backbones
C) complementary pairing of bases
D) side groups of
nitrogenous bases
A
10) Hershey and Chase set out to determine what molecule served as
the unit of inheritance. They completed a series of experiments in
which E. coli was infected by a T2 virus. Which molecular component of
the T2 virus actually ended up inside the cell?
A) protein
B) RNA
C) ribosome
D) DNA
D
11) In the polymerization of DNA, a phosphodiester bond is formed
between a phosphate group of the nucleotide being added and _____ of
the last nucleotide in the polymer.
A) the 5' phosphate
B) C6
C) the 3' OH
D) a nitrogen from the nitrogen-containing base
C
12) Replication in prokaryotes differs from replication in eukaryotes
for which of the following reasons?
A) Prokaryotic chromosomes
have histones, whereas eukaryotic chromosomes do not.
B)
Prokaryotic chromosomes have a single origin of replication, whereas
eukaryotic chromosomes have many.
C) The rate of elongation during DNA replication is slower in prokaryotes than in eukaryotes.
D) Prokaryotes produce Okazaki fragments during DNA replication, but eukaryotes do not.
B
14) Suppose you are provided with an actively dividing culture of E.
coli bacteria to which radioactive thymine has been added. What would
happen if a cell replicates once in the presence of this radioactive
base?
A) One of the daughter cells, but not the other, would have
radioactive DNA.
B) Neither of the two daughter cells would be
radioactive.
C) All four bases of the DNA would be
radioactive.
D) DNA in both daughter cells would be radioactive.
D
15) In E. coli, there is a mutation in a gene called dnaB that alters
the helicase that normally acts at the origin. Which of the following
would you expect as a result of this mutation?
A) Additional
proofreading will occur.
B) No replication fork will be formed.
C) Replication will occur via RNA polymerase alone.
D)
Replication will require a DNA template from another source.
B
16) In E. coli, which enzyme catalyzes the elongation of a new DNA
strand in the 5' → 3' direction?
A) primase
B) DNA ligase
C) DNA polymerase III
D) helicase
C
17) Eukaryotic telomeres replicate differently than the rest of the
chromosome. This is a consequence of which of the following?
A)
the evolution of telomerase enzyme
B) DNA polymerase that cannot
replicate the leading strand template to its end
C) gaps left at the 5' end of the lagging strand
D) gaps
left at the 3' end of the lagging strand because of the need for a primer
C
18) How does the enzyme telomerase meet the challenge of replicating
the ends of linear chromosomes?
A) It adds a single 5' cap
structure that resists degradation by nucleases.
B) It causes
specific double-strand DNA breaks that result in blunt ends on both
strands.
C) It catalyzes the lengthening of telomeres,
compensating for the shortening that could occur during replication
without telomerase activity.
D) It adds numerous GC pairs, which resist hydrolysis and maintain chromosome integrity.
C
19) The DNA of telomeres has been highly conserved throughout the
evolution of eukaryotes. This most likely reflects _____.
A) the
low frequency of mutations occurring in this DNA
B) continued
evolution of telomeres
C) that new mutations in telomeres have been advantageous
D) a critical function of telomeres
D
20) At a specific area of a chromosome, the sequence of nucleotides below is present where the chain opens to form a replication fork:
3' CCTAGGCTGCAATCC 5'
An RNA primer is formed starting at
the underlined T (T) of the template. Which of the following
represents the primer sequence?
A) 5' GCCTAGG 3'
B) 5'
ACGTTAGG 3'
C) 5' ACGUUAGG 3'
D) 5' GCCUAGG 3'
C
21) In E. coli, to repair a thymine dimer by nucleotide excision
repair, in which order do the necessary enzymes act?
A) nuclease,
DNA polymerase III, RNA primase
B) helicase, DNA polymerase I,
DNA ligase
C) DNA ligase, nuclease, helicase
D) nuclease, DNA
polymerase I, DNA ligase
D
22) In E. coli, what is the function of DNA polymerase III?
A) to unwind the DNA helix during replication
B) to seal
together the broken ends of DNA strands
C) to add nucleotides to
the 3' end of a growing DNA strand
D) to degrade damaged DNA molecules
C
24) The leading and the lagging strands differ in that _____.
A)
the leading strand is synthesized in the same direction as the
movement of the replication fork, and the lagging strand is
synthesized in the opposite direction
B) the leading strand is
synthesized by adding nucleotides to the 3' end of the growing strand,
and the lagging strand is synthesized by adding nucleotides to the 5'
end
C) the lagging strand is synthesized continuously, whereas
the leading strand is synthesized in short fragments that are
ultimately stitched together
D) the leading strand is synthesized
at twice the rate of the lagging strand
A
25) A new DNA strand elongates only in the 5' to 3' direction because
_____.
A) DNA polymerase begins adding nucleotides at the 5' end
of the template
B) the polarity of the DNA molecule prevents
addition of nucleotides at the 3' end
C) replication must progress toward the replication
fork
D) DNA polymerase can add nucleotides only to the free 3' end
D
27) What is the role of DNA ligase in the elongation of the lagging
strand during DNA replication?
A) It synthesizes RNA nucleotides
to make a primer.
B) It joins Okazaki fragments together.
C) It unwinds the parental double helix.
D) It stabilizes
the unwound parental DNA.
B
28) Which of the following help(s) to hold the DNA strands apart
while they are being replicated?
A) primase
B) ligase
C) DNA polymerase
D) single-strand DNA binding proteins
D
29) Individuals with the disorder xeroderma pigmentosum are
hypersensitive to sunlight. This occurs because their cells
cannot_____.
A) replicate DNA
B) undergo mitosis
C) exchange DNA with other cells
D) repair thymine dimers
D
30) Which of the following would you expect of a eukaryote lacking telomerase?
A) a high probability of somatic cells becoming
cancerous
B) an inability to produce Okazaki fragments
C)
an inability to repair thymine dimers
D) a reduction in chromosome length in gametes
D
35) Within a double-stranded DNA molecule, adenine forms hydrogen
bonds with thymine and cytosine forms hydrogen bonds with guanine.
This arrangement _____.
A) allows variable width of the double
helix
B) permits complementary base pairing
C) determines the tertiary structure of a DNA molecule
D) determines the type of protein produced
B
38) Who performed classic experiments that supported the
semiconservative model of DNA replication?
A) Watson and
Crick
B) Meselson and Stahl
C) Hershey and Chase
D) Franklin and Wilkins
B
39) DNA contains the template needed to copy itself, but it has no
catalytic activity in cells. What catalyzes the formation of
phosphodiester bonds between adjacent nucleotides in the DNA polymer
being formed?
A) ribozymes
B) DNA polymerase
C)
ATP
D) deoxyribonucleotide triphosphates
B
40) What provides the energy for the polymerization reactions in DNA synthesis?
A) ATP
B) DNA polymerase
C) breaking the hydrogen
bonds between complementary DNA strands
D) the deoxyribonucleotide triphosphate substrates
D
44) What is a telomere?
A) the mechanism that holds two sister
chromatids together
B) DNA replication during telophase
C) the site of origin
of DNA replication
D) the ends of linear chromosomes
D
45) Telomere shortening puts a limit on the number of times a cell
can divide. Research has shown that telomerase can extend the life
span of cultured human cells. How might adding telomerase affect
cellular aging?
A) Telomerase will speed up the rate of cell proliferation.
B) Telomerase eliminates telomere shortening and retards aging.
C) Telomerase shortens telomeres, which delays cellular aging.
D) Telomerase would have no effect on cellular aging.
B
46) Telomere shortening is a problem in which types of cells?
A) only prokaryotic cells
B) only eukaryotic cells
C)
cells in prokaryotes and eukaryotes
B
47) Which of the following cells have reduced or very little active telomerase activity?
A) most normal somatic cells
B) most normal germ
cells
C) most cancer cells
A
48) Researchers found E. coli that had mutation rates one hundred
times higher than normal. Which of the following is the most likely
cause of these results?
A) The single-stranded binding proteins
were malfunctioning.
B) There were one or more mismatches in the
RNA primer.
C) The proofreading mechanism of DNA polymerase was not working
properly.
D) The DNA polymerase was unable to add bases to the
end of the growing nucleic acid chain.
C
49) In a healthy cell, the rate of DNA repair is equal to the rate of
DNA mutation. When the rate of repair lags behind the rate of
mutation, what is a possible fate of the cell?
A) The cell can be
transformed to a cancerous cell.
B) RNA may be used instead of
DNA as inheritance material.
C) The cell will become embryonic.
D) DNA synthesis will
continue by a new mechanism.
A
50) Which of the following statements describes a eukaryotic
chromosome?
A) a single strand of DNA
B) a series of
nucleosomes wrapped around two DNA molecules
C) a chromosome with
different numbers of genes in different cell types of an organism
D) a single linear molecule of double-stranded DNA plus proteins
D
51) If a cell were unable to produce histone proteins, which of the
following would be a likely effect?
A) There would be an increase
in the amount of "satellite" DNA produced during centrifugation.
B) The cell's DNA couldn't be packed into its nucleus.
C) Spindle fibers would not form during prophase.
D)
Amplification of other genes would compensate for the lack of histones.
B
52) Which of the following statements is true of histones?
A)
Each nucleosome consists of two molecules of histone H1.
B)
Histone H1 is not present in the nucleosome bead; instead, it draws
the nucleosomes together.
C) The carboxyl end of each histone extends outward from the
nucleosome and is called a "histone tail."
D) Histones
are found in mammals, but not in other animals or in plants or fungi.
B
53) Why do histones bind tightly to DNA?
A) Histones are
positively charged, and DNA is negatively charged.
B) Histones are negatively charged, and DNA is positively charged.
C) Both histones and DNA are strongly hydrophobic.
D)
Histones are covalently linked to the DNA.
A
54) Which of the following represents the order of increasingly
higher levels of organization of chromatin?
A) nucleosome, 30-nm
chromatin fiber, looped domain
B) looped domain, 30-nm chromatin
fiber, nucleosome
C) nucleosome, looped domain, 30-nm chromatin fiber
D) 30-nm chromatin fiber, nucleosome, looped domain
A
56) Which of the following is most critical for the association
between histones and DNA? A) Histones are small proteins.
B)
Histones are highly conserved (that is, histones are very similar in
every eukaryote).
C) There are at least five different histone
proteins in every eukaryote.
D) Histones are positively charged.
D
57) In E. coli replication the enzyme primase is used to attach a 5
to 10 base ribonucleotide strand complementary to the parental DNA
strand. The RNA strand serves as a starting point for the
DNA
polymerase that replicates the DNA. If a mutation occurred in
the primase gene, which of the following would you expect?
A)
Replication would only occur on the leading strand.
B)
Replication would only occur on the lagging strand.
C)
Replication would not occur on either the leading or lagging
strand.
D) Replication would not be affected as the enzyme
primase in involved with RNA synthesis.
C
58) Hershey and Chase used a DNA-based virus for their work. What
would the results have been if they had used an RNA virus?
A)
With an RNA virus radioactive protein would have been in the final
pellet.
B) With an RNA virus radioactive RNA would have been in
the final pellet.
C) With an RNA virus neither sample would have had a
radioactive pellet.
D) With an RNA virus the protein shell would
have been radioactive in both samples.
B
59) The lagging strand is characterized by a series of short segments
of DNA (Okazaki fragments) that will be joined together to form a
finished lagging strand. The experiments that led to the discovery of
Okazaki
fragments gave evidence for which of the following
ideas?
A) DNA polymerase is a directional enzyme that synthesizes
leading and lagging strands during replication.
B) DNA is a
polymer consisting of four monomers: adenine, thymine, guanine, and cytosine.
C) DNA is the genetic material.
D) Bacterial replication
is fundamentally different from eukaryotic replication. The key
shouldn’t be way longer than the distractors.
A
42) What is the difference between the leading strand and the lagging strand in DNA replication?
A) The leading strand is synthesized in the 3' → 5' direction in
a discontinuous fashion, while the lagging strand is synthesized in
the 5' → 3' direction in a continuous fashion.
B) The leading
strand is synthesized continuously in the 5' → 3' direction, while
the lagging strand is synthesized discontinuously in the 5' → 3' direction.
C) The leading strand requires an RNA primer, whereas the
lagging strand does not.
D) There are different DNA polymerases
involved in elongation of the leading strand and the lagging strand.
B
43) What is a major difference between eukaryotic DNA replication and
prokaryotic DNA replication?
A) Prokaryotic replication does not
require a primer.
B) Prokaryotic chromosomes have a single origin
of replication, while eukaryotic chromosomes have multiple origins of replication.
C) DNA replication in prokaryotic cells is conservative. DNA replication in eukaryotic cells is semi-conservative.
D) DNA polymerases of prokaryotes can add nucleotides to both 3' and 5' ends of DNA strands while those of eukaryotes function only in the 5' → 3' direction.
B
3) The genetic code is essentially the same for all organisms. From
this, one can logically assume which of the following?
A) A gene
from an organism can theoretically be expressed by any other
organism.
B) DNA was the first genetic material.
C) The same codons in different organisms translate into different amino acids.
D) Different organisms have different types of amino acids.
A
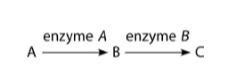
4) The figure above shows a simple metabolic pathway. According to
Beadle and Tatum's hypothesis, how many genes are necessary for this
pathway?
A) 1
B) 2
C) 3
D) It cannot be determined from the pathway.
B

5) Refer to the metabolic pathway illustrated above. If A, B, and C
are all required for growth, a strain that is mutant for the
gene-encoding enzyme A would be able to grow on medium supplemented
with _____.
A) nutrient A only
B) nutrient B only
C) nutrient C only
D) nutrients A and C
B

6) Refer to the metabolic pathway illustrated above. If A, B, and C
are all required for growth, a strain mutant for the gene-encoding
enzyme B would be able to grow on medium supplemented with
_____.
A) nutrient A only
B) nutrient B only
C) nutrient C only
D) nutrients A and C
C
10) Which of the following contradicts the one-gene, one-enzyme
hypothesis?
A) A mutation in a single gene can result in a
defective protein.
B) Alkaptonuria results when individuals lack
a single enzyme involved in the catalysis of homogentisic
acid.
C) Sickle-cell anemia results in defective
hemoglobin.
D) A single antibody gene can code for different
related proteins, depending on the splicing that takes place post-transcriptionally.
D
11) Which of the following is directly related to a single amino acid?
A) the base sequence of the tRNA
B) the amino acetyl tRNA
synthase
C) the three-base sequence of mRNA
D) the complementarity of DNA and RNA
C
12) In the process of transcription, _____.
A) DNA is replicated
B) RNA is synthesized
C) proteins
are synthesized
D) mRNA attaches to ribosomes
B
13) Codons are part of the molecular structure of _____.
A) a protein
B) mRNA
C) tRNA
D) rRNA
B
14) What does it mean when we say the genetic code is
redundant?
A) A single codon can specify the addition of more
than one amino acid.
B) The genetic code is different for different domains of
organisms.
C) The genetic code is universal (the same for all
organisms).
D) More than one codon can specify the addition of
the same amino acid.
D
15) Once researchers identified DNA as the unit of inheritance, they
asked how information was transferred from the DNA in the nucleus to
the site of protein synthesis in the cytoplasm. What is the mechanism
of information transfer in eukarotes?
A) DNA from a single gene
is replicated and transferred to the cytoplasm, where it serves as a
template for protein synthesis.
B) Messenger RNA is transcribed from a single gene and transfers
information from the DNA in the nucleus to the cytoplasm, where
protein synthesis takes place.
C) Proteins transfer information
from the nucleus to the ribosome, where protein synthesis takes place.
D) Transfer RNA takes information from DNA directly to a ribosome, where protein synthesis takes place.
B
16) According to the central dogma, what molecule should go in the
blank? DNA → _____ → Proteins
A) mtDNA
B) rRNA
C) mRNA
D) tRNA
C
17) Codons are three-base sequences that specify the addition of a
single amino acid. How do eukaryotic codons and prokaryotic codons
compare?
A) Prokaryotic codons usually contain different bases
than those of eukaryotes.
B) Prokaryotic codons usually specify
different amino acids than those of eukaryotes.
C) The translation of codons is mediated by tRNAs in eukaryotes,
but translation requires no intermediate molecules such as tRNAs in
prokaryotes.
D) Codons are a nearly universal language among all organisms.
D
18) Which of the following occurs in prokaryotes but not in eukaryotes?
A) post-transcriptional splicing
B) concurrent
transcription and translation
C) translation in the absence of a ribosome
D) gene regulation
B
19) Which of the following statements best describes the termination
of transcription in prokaryotes?
A) RNA polymerase transcribes
through the polyadenylation signal, causing proteins to associate with
the transcript and cut it free from the polymerase.
B) RNA
polymerase transcribes through the terminator sequence, causing the
polymerase to separate from the DNA and release the
transcript.
C) Once transcription has initiated, RNA polymerase
transcribes until it reaches the end of the chromosome.
D) RNA
polymerase transcribes through a stop codon, causing the polymerase to
stop advancing through the gene and release the mRNA.
B
20) In eukaryotes there are several different types of RNA
polymerase. Which type is involved in transcription of mRNA for a
globin protein?
A) RNA polymerase I
B) RNA polymerase II
C) RNA polymerase III
D) primate
B
21) Transcription in eukaryotes requires which of the following in addition to RNA polymerase?
A) start and stop codons
B) ribosomes and tRNA
C)
several transcription factors
D) aminoacyl-tRNA synthetase
C
22) Which of the following best describes the significance of the
TATA box in eukaryotic promoters?
A) It is the recognition site
for a specific transcription factor.
B) It sets the reading frame
of the mRNA.
C) It is the recognition site for ribosomal binding.
D) Its significance has not yet been determined.
A
23) Which of the following does not occur in prokaryotic gene
expression, but does occur in eukaryotic gene expression?
A)
mRNA, tRNA, and rRNA are transcribed.
B) RNA polymerase binds to
the promoter.
C) A cap is added to the 5' end of the mRNA.
D) RNA
polymerase requires a primer to elongate the molecule.
C
24) A ribozyme is _____.
A) a catalyst that uses RNA as a
substrate
B) an RNA with catalytic activity
C) an enzyme
that catalyzes the association between the large and small ribosomal subunits
D) an enzyme that synthesizes RNA as part of the transcription process
B
25) Alternative RNA splicing _____.
A) is a mechanism for
increasing the rate of translation
B) can allow the production of
proteins of different sizes and functions from a single mRNA
C) can allow the production of similar proteins from different
RNAs
D) increases the rate of transcription
B
26) In the structural organization of many eukaryotic genes,
individual exons may be related to which of the following?
A) the
sequence of the intron that immediately precedes each exon
B) the
number of polypeptides making up the functional protein
C) the various domains of the polypeptide product
D) the number of start sites for transcription
C
27) In an experimental situation, a student researcher inserts an
mRNA molecule into a eukaryotic cell after she has removed its 5' cap
and poly-A tail. Which of the following would you expect her to
find?
A) The mRNA is quickly converted into a ribosomal
subunit.
B) The cell adds a new poly-A tail to the mRNA.
C)
The mRNA attaches to a ribosome and is translated, but more
slowly.
D) The molecule is digested by enzymes because it is not
protected at the 5' end.
D
29) Which one of the following statements about RNA processing is true?
A) Exons are cut out before mRNA leaves the nucleus.
B)
Ribozymes may function in RNA splicing.
C) RNA splicing can be
catalyzed by tRNA.
D) A primary transcript is often much shorter than the final RNA molecule that leaves the nucleus.
B
30) A primary transcript in the nucleus of a eukaryotic cell is _____
the functional mRNA, while a primary transcript in a prokaryotic cell
is _____ the functional mRNA.
A) the same size as; smaller
than
B) larger than; the same size as
C) larger than; smaller than
D) the same size as; larger than
B
31) A particular triplet of bases in the coding sequence of DNA is
AAA. The anticodon on the tRNA that binds the mRNA codon is
_____.
A) TTT
B) UUA
C) UUU
D) AAA
D
32) Accuracy in the translation of mRNA into the primary structure of
a polypeptide depends on specificity in the _____.
A) binding of
ribosomes to mRNA
B) binding of the anticodon to small subunit of
the ribosome
C) attachment of amino acids to rRNAs
D) binding of the
anticodon to the codon and the attachment of amino acids to tRNAs
D
33) A mutant bacterial cell has a defective aminoacyl-tRNA synthetase
that attaches a lysine to tRNAs with the anticodon AAA instead of the
normal phenylalanine. The consequence of this for the cell will be
that _____.
A) none of the proteins in the cell will contain phenylalanine
B) proteins in the cell will include lysine instead of
phenylalanine at amino acid positions specified by the codon
UUU
C) the cell will compensate for the defect by attaching
phenylalanine to tRNAs with lysine- specifying anticodons
D) the ribosome will skip a codon every time a UUU is encountered
B
34) There are sixty-one mRNA codons that specify an amino acid, but
only forty-five tRNAs. This is best explained by the fact that
_____.
A) some tRNAs have anticodons that recognize four or more
different codons
B) the rules for base pairing between the third
base of a codon and tRNA are flexible
C) many codons are never used, so the tRNAs that recognize them are dispensable D) the DNA codes for all sixty-one tRNAs, but some are then destroyed
B
35) Which of the following is the first event to take place in
translation in eukaryotes? A) base pairing of activated
methionine-tRNA to AUG of the messenger RNA
B) binding of the
larger ribosomal subunit to smaller ribosomal subunits
C)
covalent bonding between the first two amino acids
D) the small subunit of the ribosome recognizes and attaches to the 5' cap of mRNA
D
36) A signal peptide _____.
A) directs an mRNA molecule into the
cisternal space of the ER
B) terminates translation of messenger RNA
C) helps target
a protein to the ER
D) signals the initiation of transcription
C
37) The release factor (RF) _____.
A) binds to the stop codon in
the A site in place of a tRNA
B) releases the amino acid from its
tRNA to allow the amino acid to form a peptide bond C) supplies a
source of energy for termination of translation
D) releases the
ribosome from the ER to allow polypeptides into the cytosol
A
43) What must occur before a newly made polypeptide is secreted from
a cell?
A) It must be translated by a ribosome that remains free
within the cytosol.
B) Its signal sequence must target it to the
ER, after which it goes to the Golgi.
C) Its signal sequence must
be cleaved off before the polypeptide can enter the ER.
D) Its
signal sequence must target it to the plasma membrane, where it causes exocytosis.
B
44) Translation requires _____.
A) mRNA, tRNA, DNA, and
rRNA
B) mRNA, DNA, and rRNA
C) mRNA, tRNA, and rRNA
D)
mRNA, tRNA, and DNA
C
45) During elongation, which site in the ribosome represents the
location where a codon is being read?
A) E site
B) P site
C) A site
D) the small ribosomal subunit
C
46) Once a peptide has been formed between the amino acid attached to
the tRNA in the P site and the amino acid associated with the tRNA in
the A site, what occurs next?
A) translocation
B) reading of
the next codon of mRNA
C) initiation
D) The codon-anticodon hydrogen bonds holding
the tRNA in the A site are broken.
A
47) Which one of the following, if missing, would usually prevent translation from starting?
A) exon
B) 5' cap
C) AUG codon
D) poly-A tail
C
48) Put the following events of elongation in prokaryotic translation
in chronological order. 1. Binding of mRNA with small ribosomal
subunit
2. Recognition of initiation codon
3. Complementary
base pairing between initiator codon and anticodon of initiator tRNA
4. Base pairing of the mRNA codon following the initiator codon
with its complementary tRNA 5. Attachment of the large
subunit
A) 1, 2, 3, 4, 5
B) 2, 1, 4, 3, 5
C) 5, 4, 3, 2, 1
D) 1, 2, 3, 5, 4
D
49) How does termination of translation take place?
A) The end of the mRNA molecule is reached.
B) A stop codon
is reached.
C) The 5' cap is reached.
D) The poly-A tail is reached.
B
50) Post-translational modifications of proteins may include the _____.
A) removal of introns
B) addition of a 5’ cap
C)
addition of a poly-A tail
D) addition of carbohydrates to form a glycoprotein
D
51) Which of the following statements is true about protein synthesis
in prokaryotes?
A) Extensive RNA processing is required before
prokaryotic transcripts can be translated.
B) Translation can begin while transcription is still in
progress.
C) Prokaryotic cells have complicated mechanisms for
targeting proteins to the appropriate cellular organelles.
D)
Unlike eukaryotes, prokaryotes require no initiation or elongation factors.
B
52) Which of the following types of mutation, resulting in an error
in the mRNA just after the AUG start of translation, is likely to have
the most serious effect on the polypeptide product? A) a deletion of a
codon
B) a deletion of two nucleotides
C) a substitution of the third nucleotide in an ACC codon
D) a substitution of the first nucleotide of a GGG codon
B
53) A nonsense mutation in a gene _____.
A) changes an amino
acid in the encoded protein
B) has no effect on the amino acid
sequence of the encoded protein
C) introduces a premature stop codon into the mRNA
D) alters
the reading frame of the mRNA
C
54) Which of the following DNA mutations is most likely to damage the protein it specifies?
A) a base-pair deletion
B) an addition of three
nucleotides
C) a substitution in the last base of a codon
D) a codon deletion
A
55) The most commonly occurring mutation in people with cystic
fibrosis is a deletion of a single codon. This results in
_____.
A) a base-pair substitution
B) a frameshift mutation
C) a polypeptide missing an amino acid
D) a nonsense mutation
C
56) Of the following, which is the most current description of a
gene?
A) a unit of heredity that causes formation of a phenotypic
characteristic
B) a DNA subunit that codes for a single complete
protein
C) a DNA sequence that is expressed to form a functional
product: either RNA or polypeptide
D) a discrete unit of hereditary information that consists of a sequence of amino acids
C
57) How might a single base substitution in the sequence of a gene
affect the amino acid sequence of a protein encoded by the gene, and
why?
A) Only a single amino acid could change, because the
reading frame is unaffected.
B) The amino acid sequence would be
substantially altered, because the reading frame would change with a
single base substitution.
C) All amino acids following the substitution would be affected,
because the reading frame would be shifted.
D) It is not possible
for a single base substitution to affect protein structure, because
each codon is three bases long.
A
58) An original section of DNA has the base sequence AGCGTTACCGT. A mutation in this DNA strand results in the base sequence AGGCGTTACCGT. This change represents _____.
A) a missense mutation
B) a point mutation
C) a silent mutation
D) frameshift mutation
D
59) A single base substitution mutation is least likely to be
deleterious when the base change results in _____.
A) a stop
codon
B) a codon that specifies the same amino acid as the
original codon
C) an amino acid substitution that alters the tertiary structure of the protein
D) an amino acid substitution at the active site of an enzyme
B
60) Rank the following one-base point mutations (from most likely to
least likely) with respect to their likelihood of affecting the
structure of the corresponding polypeptide.
1. insertion mutation
deep within an intron
2. substitution mutation at the third
position of an exonic codon
3. substitution mutation at the second position of an exonic codon
4. deletion mutation within the first exon of the gene
A) 1,
2, 3, 4
B) 4, 3, 2, 1
C) 2, 1, 4, 3
D) 3, 1, 4, 2
B
1) Which of the following is a protein produced by a regulatory gene?
A) operon
B) inducer
C) promoter
D) repressor
D
2) A lack of which molecule would result in a cell's inability to "turn off" genes?
A) operon
B) inducer
C) promoter
D) corepressor
D
3) Which of the following, when taken up by a cell, binds to a
repressor so that the repressor no longer binds to the
operator?
A) inducer
B) promoter
C) repressor
D) corepressor
A
4) Most repressor proteins are allosteric. Which of the following
binds with the repressor to alter its conformation?
A)
inducer
B) promoter
C) transcription factor
D) cAMP
A
5) A mutation that inactivates a regulatory gene of a repressible
operon in an E. coli cell would result in _____.
A) continuous
transcription of the structural gene controlled by that
regulator
B) complete inhibition of transcription of the
structural gene controlled by that regulator
C) irreversible binding of the repressor to the operator
D)
continuous translation of the mRNA because of alteration of its structure
A
6) The lactose operon is likely to be transcribed when _____.
A)
there is more glucose in the cell than lactose
B) there is
glucose but no lactose in the cell
C) the cyclic AMP and lactose
levels are both high within the cell
D) the cAMP level is high and the lactose level is low
C
7) Transcription of structural genes in an inducible operon _____.
A) occurs continuously in the cell
B) starts when the
pathway's substrate is present
C) starts when the pathway's
product is present
D) stops when the pathway's product is present
B
8) For a repressible operon to be transcribed, which of the following
must occur?
A) A corepressor must be present.
B) RNA
polymerase and the active repressor must be present.
C) RNA
polymerase must bind to the promoter, and the repressor must be
inactive.
D) RNA polymerase must not occupy the promoter, and the
repressor must be inactive.
C
9) Altering patterns of gene expression in prokaryotes would most
likely serve an organism's survival by _____.
A) organizing gene
expression, so that genes are expressed in a given order
B)
allowing each gene to be expressed an equal number of times
C) allowing an organism to adjust to changes in environmental conditions
D) allowing environmental changes to alter a prokaryote's genome
C
10) In positive control of several sugar-metabolism-related operons,
the catabolite activator protein (CAP) binds to DNA to stimulate
transcription. What causes an increase in CAP activity in stimulating
transcription?
A) an increase in glucose and an increase in
cAMP
B) a decrease in glucose and an increase in cAMP
C) an
increase in glucose and a decrease in cAMP
D) a decrease in
glucose and a decrease in the repressor
B
11) There is a mutation in the repressor that results in a molecule
known as a super-repressor because it represses the lac operon
permanently. Which of these would characterize such a mutant?
A)
It cannot bind to the operator.
B) It cannot make a functional repressor.
C) It cannot bind to the inducer.
D) It makes a repressor that
binds CAP.
C
Suppose an experimenter becomes proficient with a technique that allows her to move DNA sequences within a prokaryotic genome.
12) If she moves the promoter for the lac operon to the region
between the beta galactosidase (lacZ) gene and the permease (lacY)
gene, which of the following would be likely?
A) The three
structural genes will be expressed normally.
B) RNA polymerase
will no longer transcribe permease.
C) The operon will still transcribe the lacZ and lacY genes, but the mRNA will not be translated. D) Beta galactosidase will not be produced.
D
Suppose an experimenter becomes proficient with a technique that allows her to move DNA sequences within a prokaryotic genome.
13) If she moves the operator to the far end of the operon, past
the transacetylase (lacA) gene, which of the following would likely
occur when the cell is exposed to lactose?
A) The inducer will
no longer bind to the repressor.
B) The repressor will no longer
bind to the operator.
C) The operon will never be transcribed.
D) The structural
genes will be transcribed continuously.
D
Suppose an experimenter becomes proficient with a technique that allows her to move DNA sequences within a prokaryotic genome.
14) If she moves the repressor gene (lacI), along with its
promoter, to a position at some several thousand base pairs away from
its normal position, we would expect the _____.
A) repressor
will no longer bind to the operator
B) repressor will no longer
bind to the inducer
C) lac operon will be expressed continuously
D) lac operon will function normally
D
Suppose an experimenter becomes proficient with a technique that allows her to move DNA sequences within a prokaryotic genome.
15) What would occur if the repressor of an inducible operon were
mutated so that it could not bind the operator?
A) irreversible
binding of the repressor to the promoter
B) reduced transcription
of the operon's genes
C) continuous transcription of the operon's genes
D)
overproduction of catabolite activator protein (CAP)
C
Suppose an experimenter becomes proficient with a technique that allows her to move DNA sequences within a prokaryotic genome.
16) According to the lac operon model proposed by Jacob and Monod,
what is predicted to occur if the operator is removed from the
operon?
A) The lac operon would be transcribed
continuously.
B) Only lacZ would be transcribed.
C) Only lacY would be transcribed.
D) Galactosidase permease
would be produced, but would be incapable of transporting lactose.
A
7) The trp repressor blocks transcription of the trp operon when the repressor _____.
A) binds to the inducer
B) binds to tryptophan
C) is not
bound to tryptophan
D) is not bound to the operator
B
18) Extracellular glucose inhibits transcription of the lac operon by
_____.
A) strengthening the binding of the repressor to the
operator
B) weakening the binding of the repressor to the
operator
C) inhibiting RNA polymerase from opening the strands of
DNA to initiate transcription
D) reducing the levels of intracellular cAMP
D
19) CAP is said to be responsible for positive regulation of the lac operon because _____.
A) CAP binds cAMP
B) CAP binds to the CAP-binding
site
C) CAP prevents binding of the repressor to the operator
D) CAP bound to the CAP-binding site increases the frequency of transcription initiation
D
20) Imagine that you've isolated a yeast mutant that contains
histones resistant to acetylation. What phenotype do you predict for
this mutant?
A) The mutant will grow rapidly.
B) The mutant
will require galactose for growth.
C) The mutant will show low levels of gene expression.
D) The mutant will show high levels of gene expression.
C
21) The primary difference between enhancers and promoter-proximal
elements is that enhancers _____.
A) are transcription factors;
promoter-proximal elements are DNA sequences
B) enhance
transcription; promoter-proximal elements inhibit transcription
C) are at considerable distances from the promoter;
promoter-proximal elements are close to the promoter
D) are DNA
sequences; promoter-proximal elements are proteins
C
22) The reason for differences in the sets of proteins expressed in a
nerve and a pancreatic cell of the same individual is that nerve and
pancreatic cells contain different _____.
A) genes
B)
regulatory sequences
C) sets of regulatory proteins
D) promoters
C
23) Gene expression is often assayed by measuring the level of mRNA
produced from a gene. If one is interested in knowing the amount of a
final active gene product, a potential problem of this method is that
it ignores the possibility of _____.
A) chromatin condensation control
B) transcriptional control
C) alternative splicing
D) translational control
D
24) Not long ago, it was believed that a count of the number of
protein-coding genes would provide a count of the number of proteins
produced in any given eukaryotic species. This is incorrect, largely
due to the discovery of widespread _____.
A) chromatin
condensation control
B) transcriptional control
C) alternative splicing
D) translational control
C
25) One way to detect alternative splicing of transcripts from a
given gene is to _____.
A) compare the DNA sequence of the given
gene to that of a similar gene in a related organism
B) measure the relative rates of transcription of the given gene
compared to that of a gene known to be constitutively spliced
C)
compare the sequences of different primary transcripts made from the
given gene
D) compare the sequences of different mRNAs made from
the given gene
D
26) Which of the following mechanisms is (are) used to coordinate the
expression of multiple, related genes in eukaryotic cells?
A)
Environmental signals enter the cell and bind directly to
promoters.
B) The genes share a single common enhancer, which
allows appropriate activators to turn on their transcription at the
same time.
C) The genes are organized into a large operon, allowing them to be
coordinately controlled as a single unit.
D) A single repressor
is able to turn off several related genes.
B
27) DNA methylation and histone acetylation are examples of _____.
A) genetic mutation
B) chromosomal rearrangements
C)
epigenetic phenomena
D) translocation
C
28) In eukaryotes, general transcription factors _____
A) bind
to other proteins or to the TATA box
B) inhibit RNA polymerase
binding to the promoter and begin transcribing
C) usually lead to
a high level of transcription even without additional specific
transcription factors
D) bind to sequences just after the start
site of transcription
A
30) Which of the following is most likely to have a small protein called ubiquitin attached to it?
A) a cyclin protein, that usually acts in G1, in a cell that is in
G2
B) a cell surface protein that requires transport from the
ER
C) an mRNA leaving the nucleus to be translated
D) an mRNA produced by an egg cell that will be retained until after fertilization
A
A researcher found a method she could use to manipulate and quantify phosphorylation and methylation in embryonic cells in culture.
31) In one set of experiments she succeeded in increasing
acetlylation of histone tails. Which of the following results would
she most likely see?
A) increased chromatin condensation
B)
decreased chromatin condensation
C) decreased binding of transcription factors
D) inactivation of the selected genes
B
A researcher found a method she could use to manipulate and quantify phosphorylation and methylation in embryonic cells in culture.
32) One of her colleagues suggested she try increased methylation of C nucleotides in the DNA of promoters of a mammalian system. Which of the following results would she most likely see?
A) decreased chromatin condensation
B) activation of histone
tails for enzymatic function
C) higher levels of transcription of certain genes D) inactivation of the selected genes
D
A researcher found a method she could use to manipulate and quantify phosphorylation and methylation in embryonic cells in culture.
33) Which method is utilized by eukaryotes to control their gene
expression that is NOT used in bacteria?
A) control of chromatin
remodeling
B) control of RNA splicing
C) transcriptional control
D) control of both RNA splicing and
chromatin remodeling
D
34) The phenomenon in which RNA molecules in a cell are destroyed if
they have a sequence complementary to an introduced double-stranded
RNA is called _____.
A) RNA interference
B) RNA obstruction
C) RNA blocking
D) RNA disposal
A
35) At the beginning of this century there was a general announcement regarding the sequencing of the human genome and the genomes of many other multicellular eukaryotes. Many people were surprised that the number of protein-coding sequences was much smaller than they had expected. Which of the following could account for much of the DNA that is not coding for proteins?
A) DNA that consists of histone coding sequences
B) DNA that is
translated directly without being transcribed
C)
non-protein-coding DNA that is transcribed into several kinds of small
RNAs with biological function
D) non-protein-coding DNA that
serves as binding sites for reverse transcriptase
C
37) Which of the following best describes siRNA?
A) a
double-stranded RNA, one of whose strands can complement and
inactivate a sequence of mRNA
B) a single-stranded RNA that can,
where it has internal complementary base pairs, fold into cloverleaf
patterns
C) a double-stranded RNA that is formed by cleavage of
hairpin loops in a larger precursor
D) a portion of rRNA that
allows it to bind to several ribosomal proteins in forming large or
small subunits
A
A researcher introduces double-stranded RNA into a culture of mammalian cells and can identify its location or that of its smaller subsections experimentally, using a fluorescent probe.
38) Some time later, she finds that the introduced strand separates
into single-stranded RNAs, one of which is degraded. What does this
enable the remaining strand to do?
A) attach to histones in the
chromatin
B) bind to complementary regions of target mRNAs
C) activate other siRNAs in the cell
D) bind to
noncomplementary RNA sequences
B
A researcher introduces double-stranded RNA into a culture of mammalian cells and can identify its location or that of its smaller subsections experimentally, using a fluorescent probe.
39) When she finds that the introduced strand separates into
single-stranded RNAs, what other evidence of this single-stranded RNA
piece's activity can she find?
A) She can measure the degradation
rate of the remaining single strand.
B) The rate of accumulation
of the polypeptide encoded by the target mRNA is reduced.
C) The amount of miRNA is multiplied by its replication.
D) The cell's translation ability is entirely shut down.
B
47) In colorectal cancer, several genes must be mutated for a cell to
develop into a cancer cell. Which of the following kinds of genes
would you expect to be mutated?
A) genes coding for enzymes that
act in the colon
B) genes involved in control of the cell cycle
C) genes that are especially susceptible to mutation
D) genes
of the bacteria, which are abundant in the colon
B
51) Which of the following types of mutation would convert a proto-oncogene into an oncogene?
A) a mutation that blocks transcription of the
proto-oncogene
B) a mutation that creates an unstable
proto-oncogene mRNA
C) a mutation that greatly increases the
amount of the proto-oncogene protein
D) a deletion of most of the proto-oncogene coding sequence
C
52) Proto-oncogenes _____.
A) normally suppress tumor
growth
B) are produced by somatic mutations induced by
carcinogenic substances
C) stimulate normal cell growth and division
D) are
underexpressed in cancer cells
C
53) The product of the p53 gene _____.
A) inhibits the cell
cycle
B) slows down the rate of DNA replication by interfering
with the binding of DNA polymerase C) causes cells to reduce
expression of genes involved in DNA repair
D) allows cells to
pass on mutations due to DNA damage
A
54) Tumor-suppressor genes _____.
A) are frequently
overexpressed in cancerous cells
B) are cancer-causing genes
introduced into cells by viruses
C) encode proteins that help prevent uncontrolled cell growth
D) often encode proteins that stimulate the cell cycle
C
56) Forms of the Ras protein found in tumors usually cause which of the following?
A) DNA replication to stop
B) cell-to-cell adhesion to be
nonfunctional
C) cell division to cease
D) excessive cell division
D